Gladstone NOW: The Campaign Join Us on the Journey✕
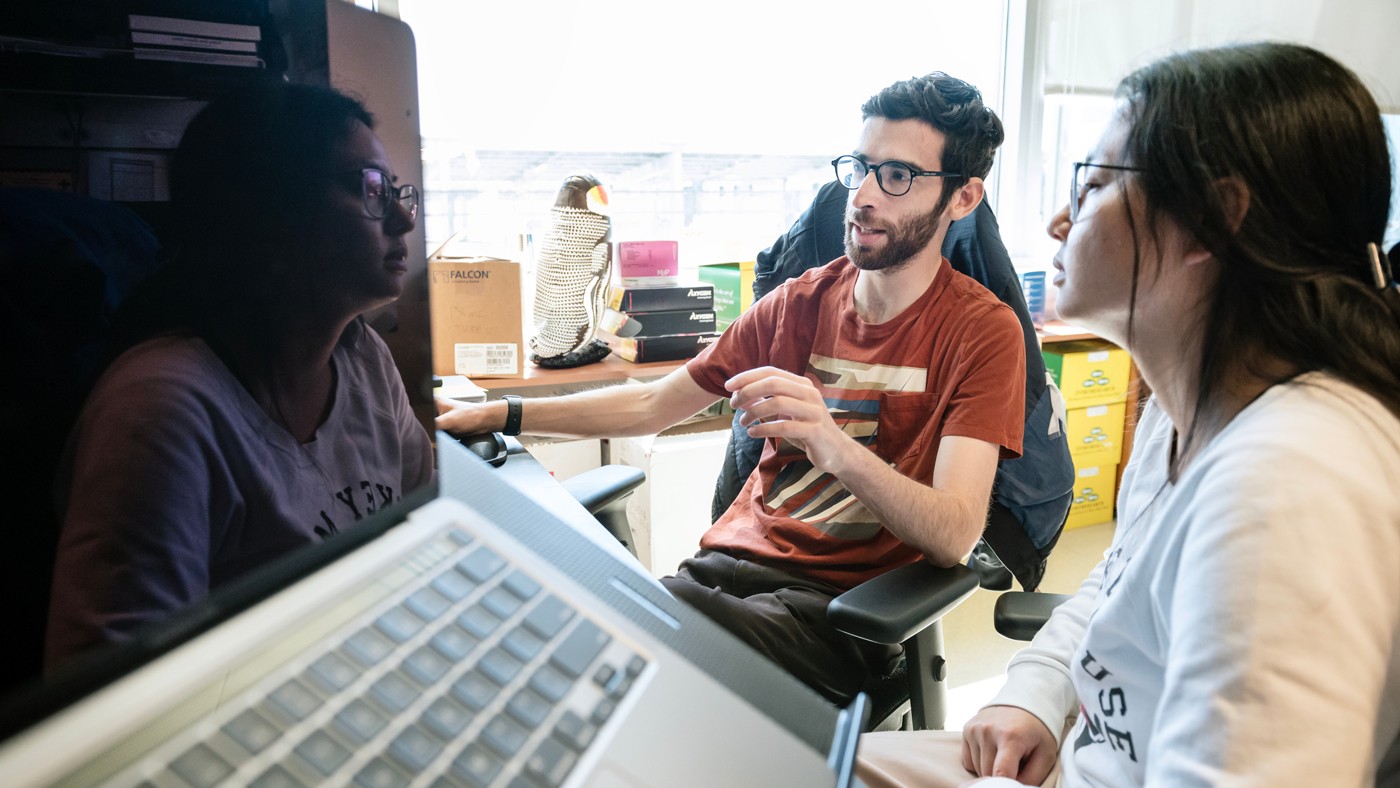

Scientists in Jennifer Doudna's lab—including Jason Nomburg (left) and Karen Zhu (right)—provide novel insights into nearly 70,000 lesser-known viral proteins that could eventually help in the development of new antiviral therapies.
Viruses are tricky to keep up with. They evolve quickly and regularly develop new proteins that help them infect their hosts.
These rapid shifts mean that researchers are still trying to understand a multitude of viral proteins and precisely how they increase viruses’ infecting abilities—knowledge that could be crucial for developing new or better virus-fighting treatments.
Now, a team of scientists at Gladstone Institutes and the Innovative Genomics Institute led by Jennifer Doudna, PhD, have harnessed computational tools to predict the three-dimensional shapes of nearly 70,000 viral proteins.
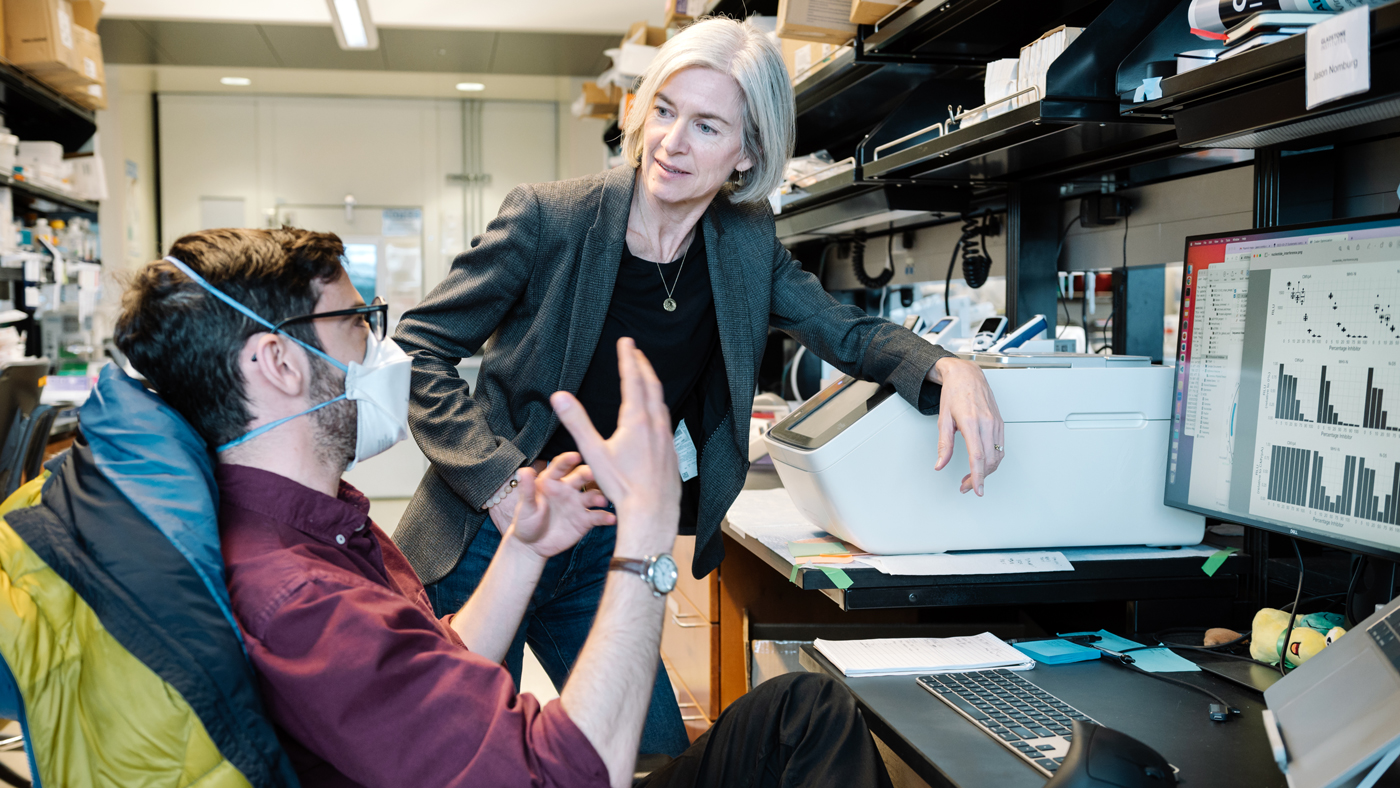

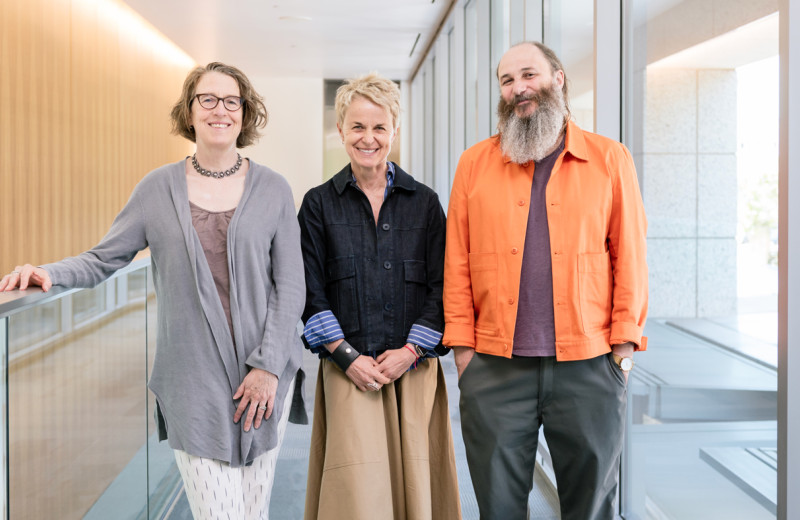
In a new study, Jason Nomburg (left) and Jennifer Doudna (right) uncovered an ancient immune-evading strategy shared by animal viruses and viruses that infect bacteria.
The researchers matched the 3D shapes to the structures of proteins whose functions are already known. Because the structure of a protein directly contributes to its biological function, their study provides new insights into what, exactly, these lesser-known proteins do.
Among their other findings, published in the journal Nature, the researchers discovered a powerful way that viruses evade immune systems. In fact, they found that bacteria-infecting viruses and those that infect higher organisms—including humans—share a similar, ancient mechanism to evade host immune defenses.
“As viruses with pandemic potential emerge, it’s important to establish how they’ll interact with human cells,” says Doudna, who is also a professor at UC Berkeley and a Howard Hughes Medical Institute Investigator. “Our new study provides a tool to predict what those newly emerging viruses can do.”
Sequence Versus Shape
Typically, to figure out the function of a protein, researchers will look for similarities between its distinct sequence of amino-acid “building blocks” and the amino-acid sequences of other proteins with known functions. However, because viruses evolve so rapidly, many viral proteins lack strong similarities with known proteins.
Still, just as different combinations of building blocks can be used to construct very similar structures, proteins with different sequences may share 3D shapes and play similar biological roles.
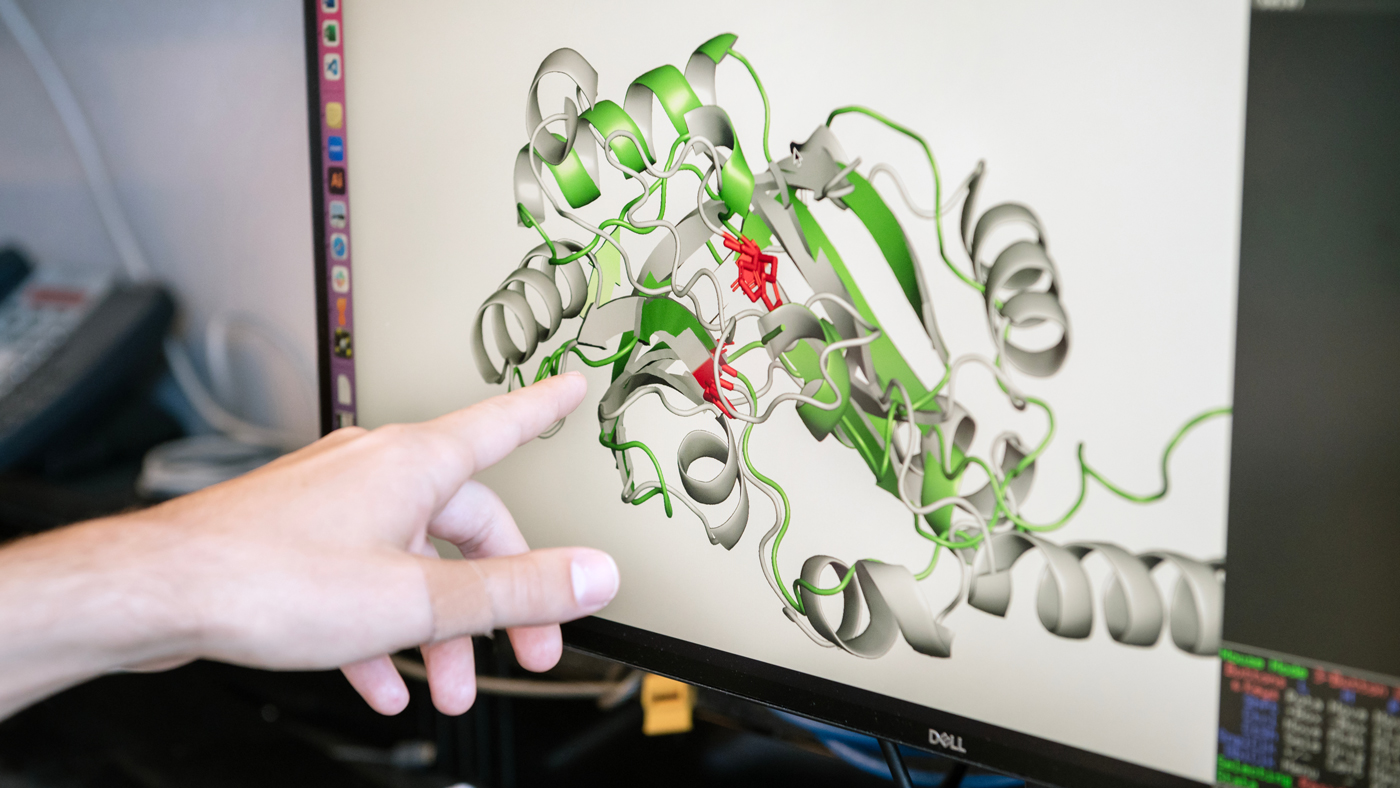
Nomburg and his colleagues used computational and deep learning tools to predict the 3D shapes of proteins from nearly 4,500 species of viruses that infect animals and humans, providing clues to their biological functions.
“We looked at similarities between protein shapes as a promising alternative for determining the function of viral proteins,” says Jason Nomburg, PhD, a postdoctoral scholar in Doudna’s lab at Gladstone and first author of the study. “We asked: what can we learn from protein structures that we might miss when just considering sequences?”
To answer that question, the team turned to an open-access research platform called AlphaFold, which predicts the 3D shape of a protein based on its amino-acid sequence. They used AlphaFold to predict the shapes of 67,715 proteins from nearly 4,500 species of viruses that infect eukaryotes (organisms including plants, animals, and humans that contain DNA in their cells’ nucleus). Then, using a deep learning tool, they compared the predicted structures with those of known proteins from other viruses, as well as non-viral proteins from eukaryotes.
“This would not have been possible without recent advancements in these types of computational tools that allow us to accurately and quickly predict and compare protein structures,” Nomburg says.
Unexpected Connections
The team discovered that 38 percent of the newly predicted protein shapes matched previously known proteins and found key connections between them.
For instance, some of the newly predicted structures belong to the group of so-called “UL43-like proteins,” which are found in human herpesviruses, including those causing mononucleosis and chicken pox.
“These new viral proteins look shockingly similar to known non-viral proteins in mammalian cells that help transport the building blocks of DNA and RNA across membranes,” Nomburg says. “Prior to this work, we didn’t know that these proteins may function as transporters.”
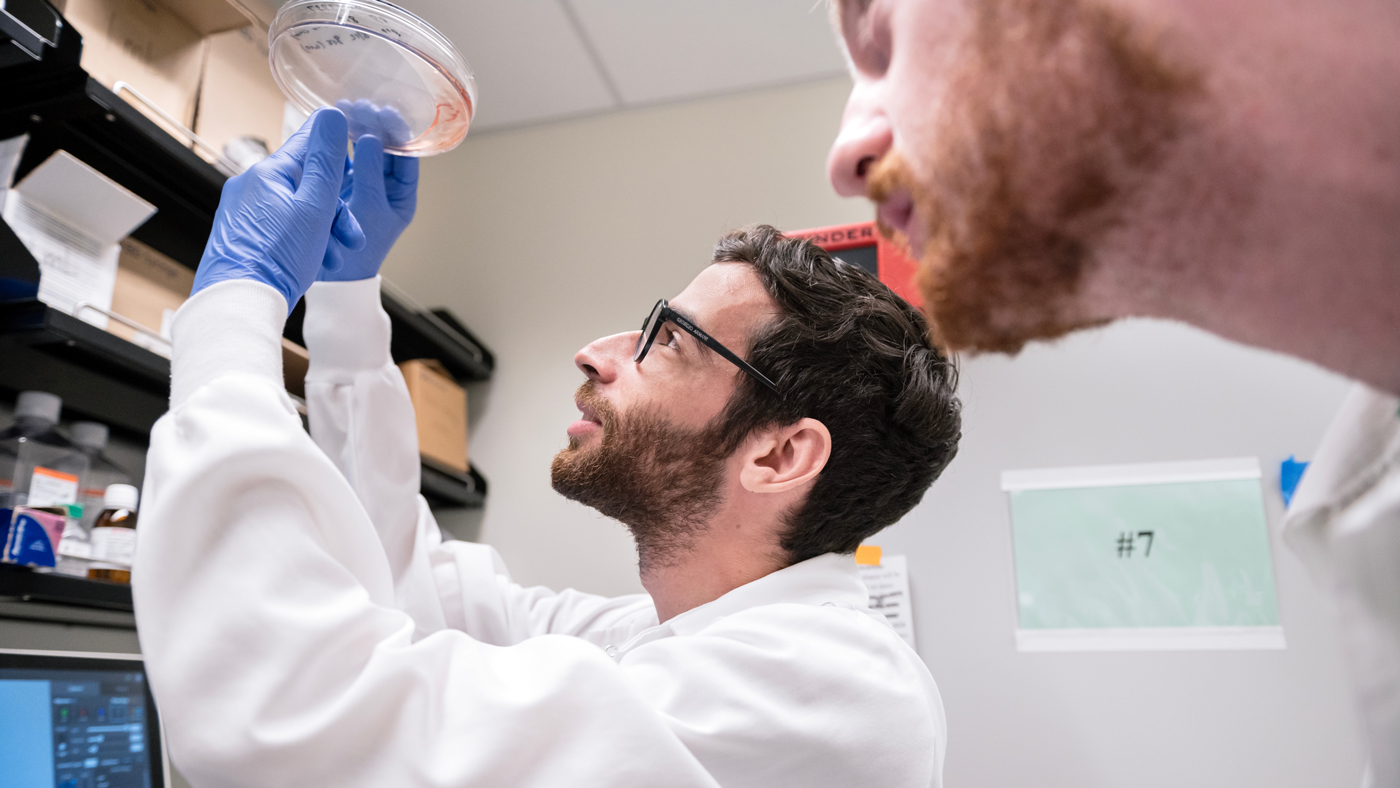

The team's findings and data analysis is publicly available, so researchers worldwide can continue to uncover how viruses interact with their hosts. Seen here are Jason Nomburg (left) and Nate Price (right).
The team also found matches between the newly predicted viral protein structures and the structures of other viral proteins. Most notably, the analysis revealed a strategy for evading host immune defenses that is broadly shared across viruses that infect animals and viruses known as phages that infect bacteria. This mechanism appears to have been conserved throughout the course of evolution.
“This is getting into a very exciting area because there is growing evidence that the innate immunity of complex organisms, including humans, resembles many different types of innate immunity in bacteria,” Nomburg says. “We will be looking more deeply into these evolutionary connections, because a better understanding of the ways our cells respond to viruses could lead to new approaches for enhancing antiviral defenses.”
Meanwhile, the team has made the 70,000 newly predicted viral protein structures, as well as data from their new analyses, publicly available. These resources could provide other researchers with opportunities to discover additional structural connections between proteins that deepen knowledge of how viruses interact with their hosts.
“From the perspective of tackling disease, this work is exciting because it highlights new possible approaches for designing broadly effective antiviral therapies,” says Doudna. “For instance, finding common, conserved ways that viruses evade immunity could lead to potent antivirals that are effective against many different viruses at once.”
For Media
Julie Langelier
Associate Director, Communications
415.734.5000
Email
About the Study
The paper “Birth of protein folds and functions in the virome” was published in the journal Nature on August 26, 2024. The authors are Jason Nomburg, Nathan Price, Yong K. Zhu, and Jennifer Doudna of Gladstone Institutes, UC Berkeley, and the Innovative Genomics Institute; and Erin E. Doherty and Daniel Bellieny-Rabelo of UC Berkeley and the Innovative Genomics Institute.
The work was supported by the Howard Hughes Medical Institute, the National Institutes of Health (U19AI171110, U54AI170792, U19AI135990, UH3AI150552, U01AI142817, U19NS132303, R21HL173710, and F32GM153031), the National Science Foundation (2334028), the Department of Energy (DE-AC02-05CH11231, 2553571 and B656358), Lawrence Livermore National Laboratory, Apple Tree Partners (24180), the UCB-Hampton University Summer Program, L. K. Shing, Koret-Berkeley-TAU, the Emerson Collective, and the Innovative Genomics Institute.
About Gladstone Institutes
Gladstone Institutes is an independent, nonprofit life science research organization that uses visionary science and technology to overcome disease. Established in 1979, it is located in the epicenter of biomedical and technological innovation, in the Mission Bay neighborhood of San Francisco. Gladstone has created a research model that disrupts how science is done, funds big ideas, and attracts the brightest minds.
Featured Experts
Gladstone Launches Center for PhAIge Therapy to Harness AI in the Fight Against Drug-Resistant Infections
Gladstone Launches Center for PhAIge Therapy to Harness AI in the Fight Against Drug-Resistant Infections
The center, funded by an NIH grant, will become one of three national centers dedicated to accelerating the development of phage therapy.
News Release Grants Data Science and Biotechnology Infectious Disease Shipman Lab Pollard Lab Ott Lab Silas Lab AIGladstone Investigator Andrew Yang Receives 2026 Sobrato Prize in Neuroscience
Gladstone Investigator Andrew Yang Receives 2026 Sobrato Prize in Neuroscience
The prize honors scientists whose breakthroughs in brain research have high potential for patient impact.
Philanthropy Awards News Release Alzheimer’s Disease Neurological Disease Yang LabMapping How the Brain Takes Out Its Trash
Mapping How the Brain Takes Out Its Trash
Gladstone scientists developed a new tool to trace how the brain clears “waste,” uncovering surprising new biology about how it keeps itself healthy.
News Release Research (Publication) Alzheimer’s Disease Neurological Disease Yang Lab




