Gladstone NOW: The Campaign Join Us on the Journey✕

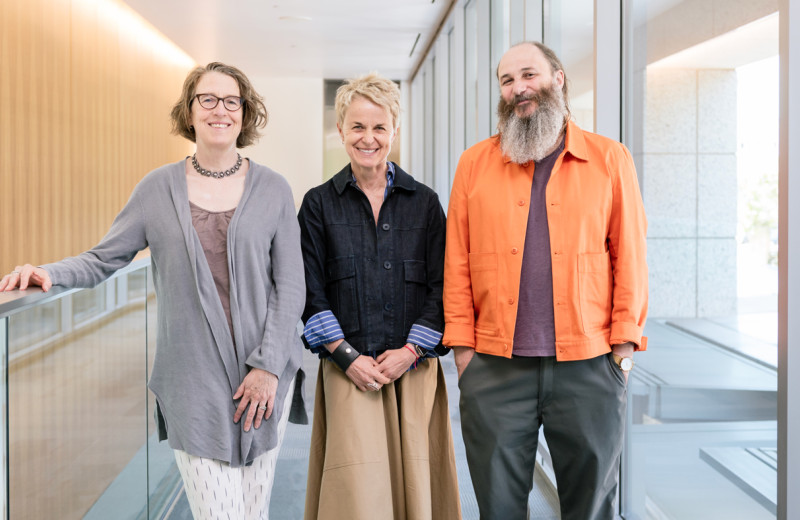
Gladstone scientists who develop new technologies—both AI models on the computer and molecular tools in the lab—are coming together to solve one of the most challenging problems in science: what specific changes in the human genome cause disease?
This article is part of a series about the many ways our scientists are using—and developing—AI tools for biomedical research. Sign up for our newsletter to have these stories delivered right to your inbox.
Your genome is made up of all the DNA you inherit—about 3 billion letters of biological code. Today, you can have your genome sequenced for a few hundred dollars. That means you can know the exact order of all those DNA letters, a blueprint which is unique to you.
Although your DNA is about 99.9 percent the same as all other humans, the remaining tiny fraction contains millions of small differences (called genetic variants). These differences can influence how your body functions, how you respond to medications, and your chances of developing certain diseases.
“The ultimate goal would be to go see your doctor, have your genome sequenced, and have your doctor tell you what your specific genetic variants mean, whether they’re related to the symptoms you’re experiencing, and whether they might increase your risk for a disease,” says Seth Shipman, PhD, investigator at Gladstone Institutes.
But we’re not there just yet. Scientists and clinicians currently don’t know how to fully decipher the genome and figure out which of the 3 billion letters matter.
“We’re about to enter a world where everybody will have their DNA sequenced, but most of the information we get from a genome sequence is still not interpretable,” says Deepak Srivastava, MD, president of Gladstone. “Only a few of the millions of genetic changes are meaningful for disease, and we can’t yet pinpoint what those are.”
So, why is it so difficult to identify the exact genetic cause of disease? In part, it’s because such a huge number of DNA changes could be responsible that scientists can’t test all the possibilities. Without the ability to prioritize the countless options, progress has been slow, or even stalled.
This is where artificial intelligence, or AI, can completely shift the paradigm.
“By using AI to predict outcomes before ever going into a lab, we can increase the speed of discovery a millionfold.”
—Katie Pollard, PhD
“We can run thousands of experiments on the computer in one day that would take years in a traditional lab,” says Katie Pollard, PhD, director of the Gladstone Institute of Data Science and Biotechnology. “By using AI to predict outcomes before ever going into a lab, we can increase the speed of discovery a millionfold.”
Pollard and her colleagues are set on leveraging the power of AI to decode the entire genome and, for the first time, actually figure out what it means.
“We’re essentially trying to establish a lookup table,” says Gladstone Investigator Vijay Ramani, PhD. “If a patient comes in, we could then find their genetic variant and let them know how likely they are to develop a particular disease.”
Where AI Meets the Lab
At Gladstone, scientists who develop new technologies—both on the computer and in the lab—have joined forces to build a platform that combines advanced computational models with novel tools in the experimental lab to test their hypotheses about how DNA works.
The first component of this platform is an AI model, designed by Pollard’s team, that can look at millions of genome sequences and make predictions about which DNA changes are most likely causing disease.
These AI predictions won’t all be correct, so they are then tested in the Shipman Lab. This group invented a new genome editing method to edit the DNA of hundreds or thousands of human cells simultaneously in a single dish.
Then, the edited cells are analyzed using a new measurement technology developed by Ramani, which can confirm the edit that each cell received and, importantly, measure the effect of each edit on the function of the cell.
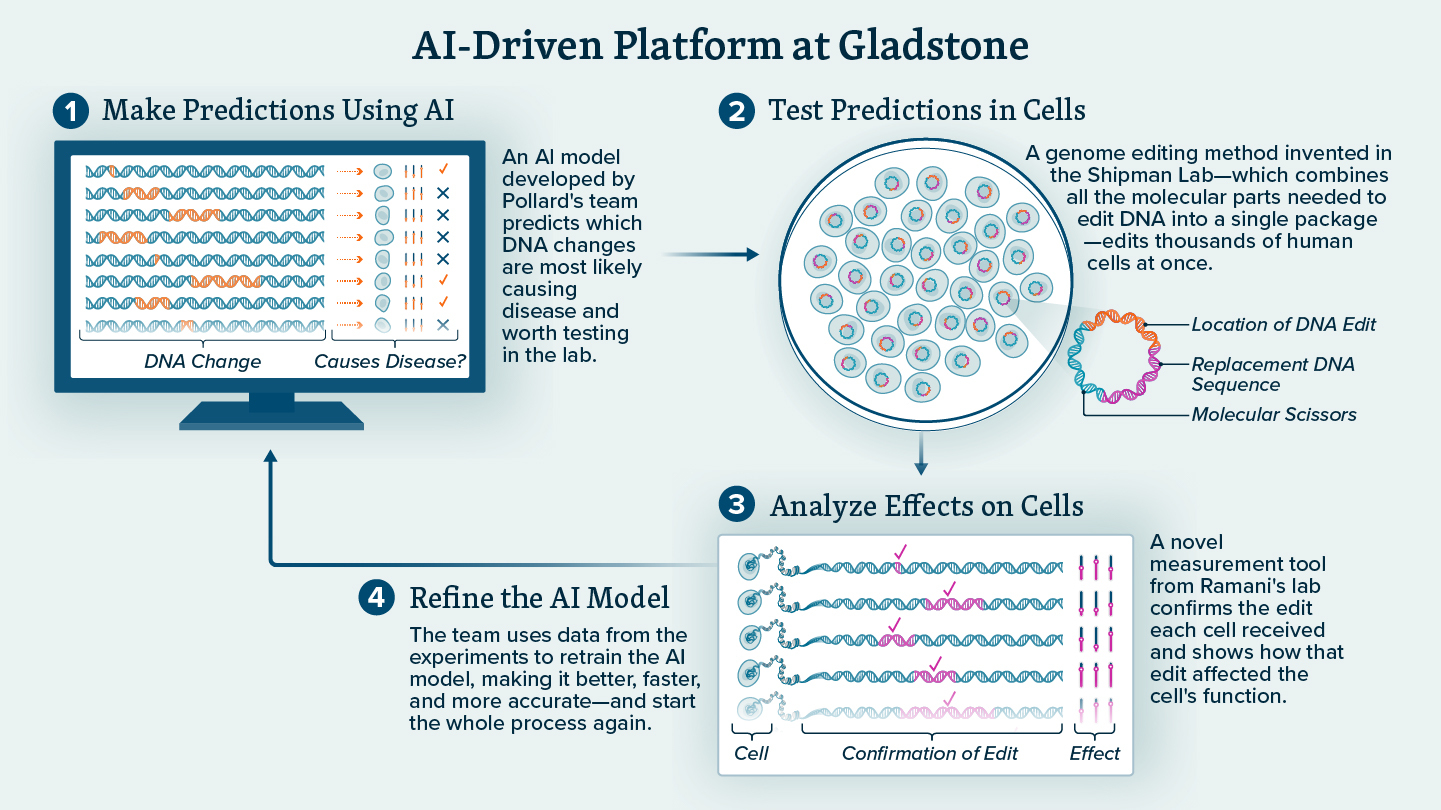
Finally, the scientists use all the data from the experiments to retrain the AI model, telling it which predictions were right and which were wrong, to make it better, faster, and more accurate—and start the whole process again.
“Nobody has tried this at this scale before,” Pollard says. “It’s been possible to make a single change, or a small number of changes, in DNA, but our team is going to interrogate the entire genome and try to understand how each letter of the genome works.”
This joint solution didn’t come about easily. Pollard, Shipman, and Ramani have had continuous discussions for over a year to get to this point.
“The fact that our labs can collaborate so seamlessly has been crucial,” Ramani says. “We kept going back and forth to figure out how we could improve our own technologies in a way that would overcome the bottlenecks in our fields. By working toward this common goal, we each pushed the limits of our labs and we can now achieve something unprecedented.”
A Quick Lesson in Genomics
Scientists used to assume disease happens when a gene is broken. So when they first sequenced the human genome in the early 2000s, they thought they would finally understand what causes disease. Instead, they found something unexpected: most disease-linked genetic differences aren’t actually in genes that make proteins—they’re in non-coding DNA.
Non-coding DNA, which makes up 98–99 percent of the genome, acts as a control panel, providing instructions for when, where, and how strongly genes are used. If you think of genes as lights in a skyscraper, non-coding DNA would be the switches and dimmers that control them. So, as it turns out, disease is most often not caused by broken bulbs, but faulty switches—perhaps causing a light to turn on when it shouldn’t or making it shine too dimly.
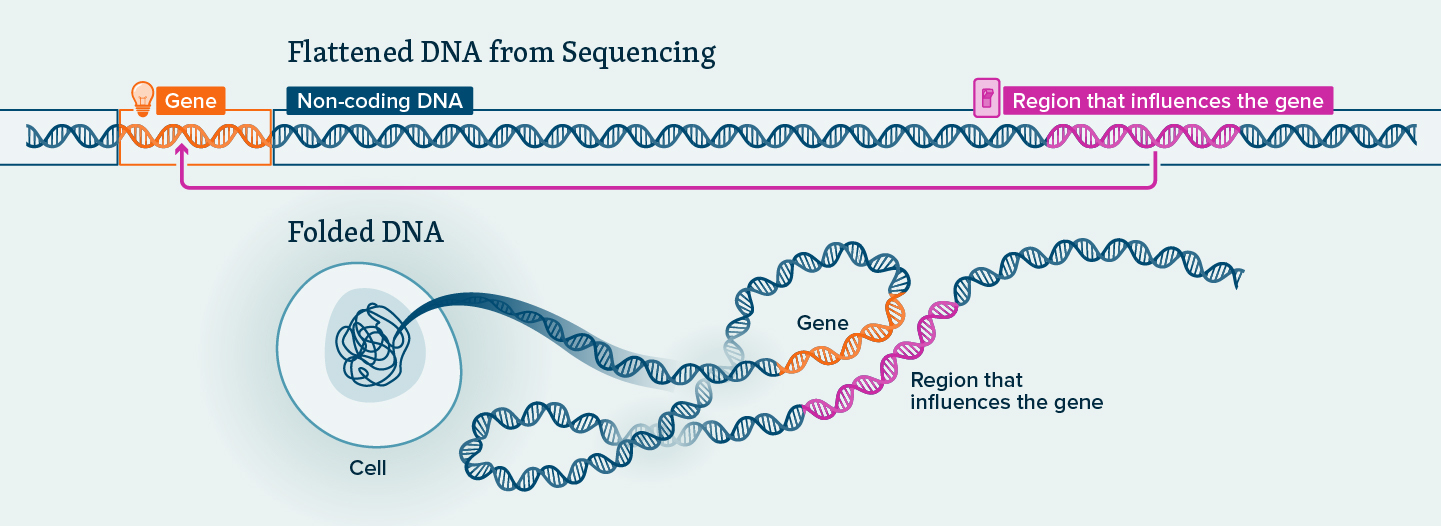
Studying non-coding DNA has been one of the hardest problems in genomics.
Its scale is overwhelming, and regions may only be active in one specific cell type, making it difficult to replicate in the lab. Still, the biggest challenge is the way DNA is neatly folded in 3D space to fit inside the nucleus of a cell. A stretch of non-coding DNA can influence a gene that’s hundreds of thousands of letters away. When DNA is folded, these two sections become physically close to one another. But with traditional sequencing techniques, which flatten DNA into a linear readout, those important spatial relationships become invisible.
These are exactly the types of problems AI is designed for—finding subtle patterns across enormous, complex datasets that humans would never spot.
From a Million Possibilities to a Thousand
Gladstone’s revolutionary platform begins in Pollard’s lab, where she and her team develop AI models using deep learning. Deep learning is a way for a computer to recognize patterns by learning on its own from a huge number of examples, and then use those patterns to make predictions about data it has never seen.
This AI approach is called “deep” because it uses many layers of processing. Each layer performs a simple operation, but when they’re combined, the system can learn very complex patterns—much like each step of a factory assembly line does a simple task, but at the end, the tasks can result in a very elaborate product.
“That’s where AI modeling comes in, because we can do trillions of experiments on the computer and predict the ones worth testing in the lab.”
—Katie Pollard, PhD
“Even with the advanced experimental tools my colleagues are building, it’s still not possible to test every theory we have about how DNA works,” Pollard says. “That’s where AI modeling comes in, because we can do trillions of experiments on the computer and predict the ones worth testing in the lab.”
Pollard’s predictive model was trained using millions of DNA sequences, from both healthy individuals and patients with disease. Now, the scientists can input a DNA sequence and the AI model will predict how the cells will function. It can forecast which genes are turned on or off, how the genome is folded in 3D space, and which stretches of non-coding DNA are important. This is especially valuable for rare diseases where patient samples for functional experiments are hard to come by, but DNA sequencing can be done cheaply using only saliva or blood.
The researchers can also probe the model to find out how it reaches its conclusions. By asking AI to show its work, the team can learn new insights into how DNA instructs cells to behave.
“Our model has learned the rules that are encoded in DNA,” Pollard says. “After years of plodding along, I feel like with these AI models, we’re suddenly moving at lightning speed and discoveries are coming so much faster than before.”
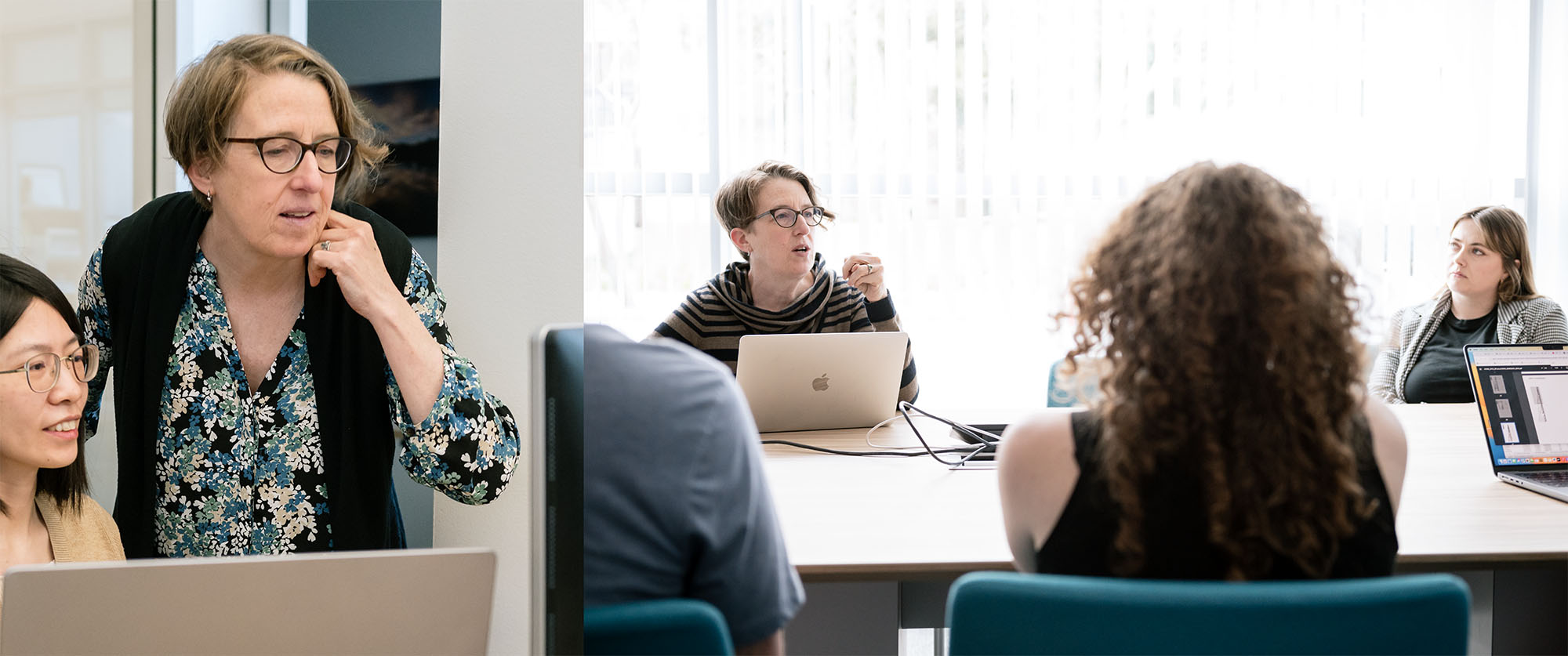

Katie Pollard (right, in the photo on the left; and center in the other photo) and her team have built an AI model that has learned the rules encoded in DNA and can now predict what would happen to a cell if a stretch of DNA is changed. (In the photo on the left, Pollard is seen with Bioinformatics Fellow Zhirui Hu.)
The team can now conduct genome editing experiments on the computer. In a matter of seconds, they can change a particular stretch of DNA, say by deleting a sequence that the model thinks is important even though it has never been studied in the lab, and the model will correctly predict what would happen to the cell.
Take congenital heart defects: millions of DNA letters could be contributing to this disease. Pollard’s lab can use AI to prioritize those options and narrow them down to a few thousand most likely to be involved.
“We can get it down by two whole orders of magnitude, from a million to a thousand,” Pollard explains. “But verifying each of those predictions in the lab, using a traditional genome editing approach, would take about two months per experiment. You just can’t do a thousand experiments like that consecutively.”
Enter the Shipman Lab. They created a new technique to edit the DNA of thousands of cells at once—and the entire process only takes one or two days.
Thousands of Edits, One Experiment
Genome editing is a way to make targeted changes to DNA inside a living cell—removing, adding, or replacing sequences of DNA letters at a precise location. By changing the genome and analyzing the effect of that change, scientists can understand how cells work and how they break in disease.
In 2012, Gladstone Investigator Jennifer Doudna, PhD, co-discovered a now-famous genome editing technique known as CRISPR, which made the technology more accessible and affordable for labs around the world.
Here’s how it usually works. First, scientists assemble molecular scissors that can cut the DNA and a targeting molecule that guides them, much like a GPS system, to the right spot on the genome. To insert new DNA into the genome, the scientists also need a replacement sequence, called “template DNA.” This is often delivered separately to the cell.
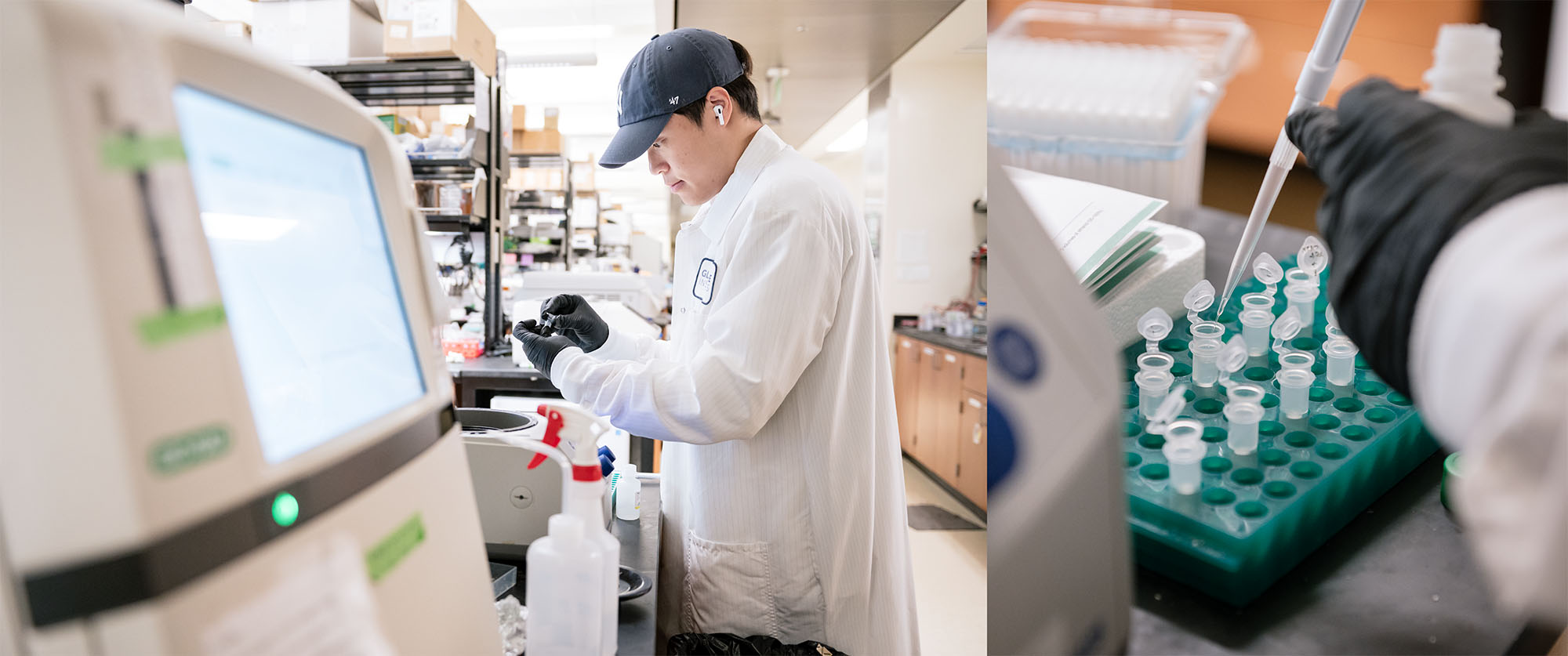

The technology developed in the Shipman Lab reduces the time needed to edit DNA in cells from six weeks to two days. (Seen here is Jihoon Han, postdoctoral scholar in Shipman’s lab.)
Once the genome edit is done, scientists then go through a cloning step. This involves separating the cells and letting them grow into individual colonies. Finally, to confirm the cells received the correct edit, scientists need to extract a small piece of DNA from each colony and sequence it—reading out the DNA letters to ensure the desired change is there.
This entire process usually takes about six weeks. If a researcher wanted to test many different DNA edits, they would need to repeat this process for each one.
“Now, we can effectively do one single experiment that actually tests thousands of genome edits simultaneously.”
—Seth Shipman, PhD
“We found a way to simplify and speed up the process,” says Shipman. “We can package all the molecular parts needed for genome editing into a single package, and deliver them in bulk to a dish of cells, so that every cell in the dish gets a different edit.”
Shipman and his team use retrons, which are bacterial immune systems that act like miniature DNA factories. They pioneered a technique that combines elements from the CRISPR system with retrons, and bundles them into one small circular piece of DNA, called a plasmid. Each plasmid can contain instructions for a unique gene edit.
When the plasmid gets delivered to a cell, the molecular scissors cut the DNA, and the retrons produce template DNA directly inside a cell, providing the necessary genetic material for repair.
“Having all the editing components self-contained on a plasmid is critical,” Shipman says. “This system enables us to make hundreds or thousands of edits at the same time in a pool of cells, while making sure that each cell gets everything it needs to carry out a specific edit, without interference.”
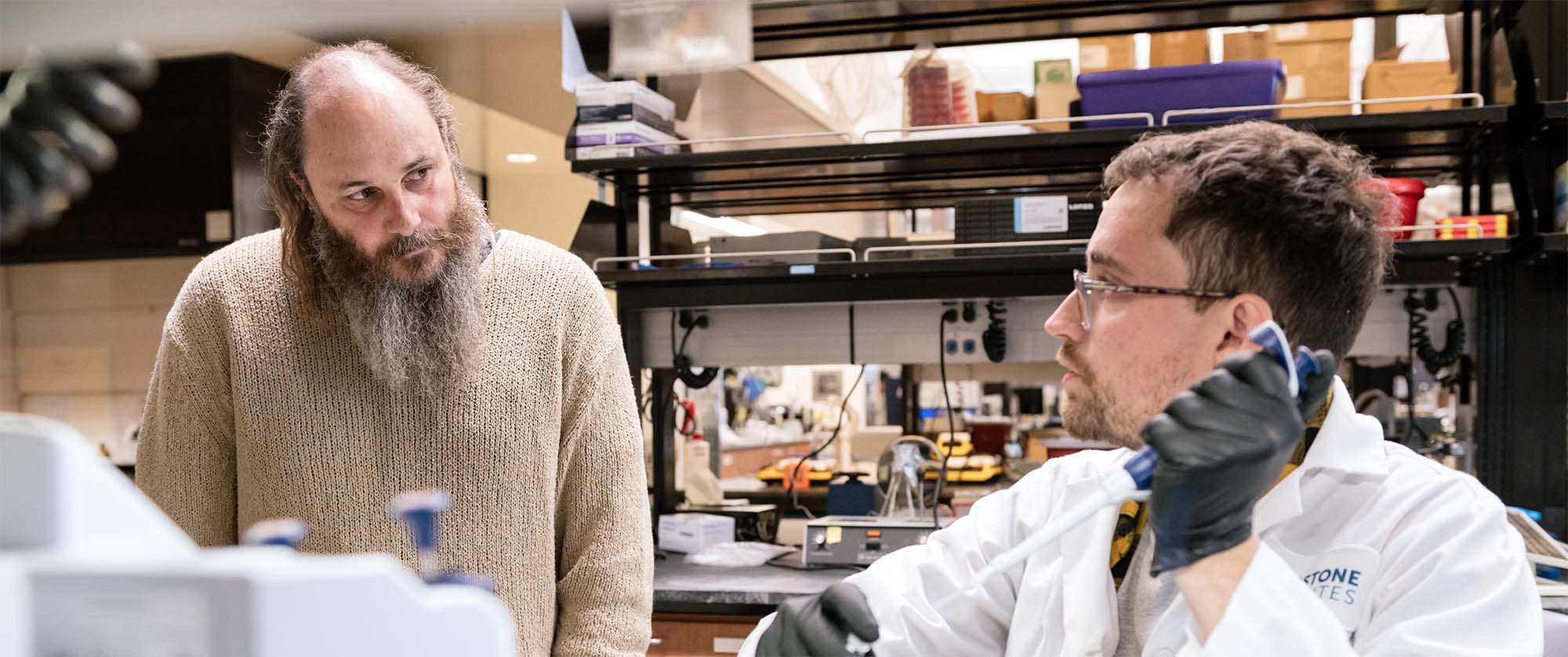

Seth Shipman (left), seen here in the lab with Alejandro González-Delgado (right), invented a way to make thousands of genome edits simultaneously—in one single experiment.
In contrast, when template DNA is delivered to cells separately from the editing materials, things could go wrong. Some cells could end up with the wrong combination of molecular scissor and replacement DNA, or only receive one or the other.
“Our goal is to avoid having to conduct many experiments one by one, and instead design a way to test the thousands of predictions made by the AI model at once,” Shipman says. “Now, we can effectively do one single experiment that actually tests thousands of genome edits simultaneously.”
Shipman also found a way to skip the time-consuming cloning step, using a novel measurement technique developed by the Ramani Lab. Rather than having to separate the cells and confirm each one got the correct edit, they can analyze all the cells at once and not only figure out which cell received which edit, but also determine how that edit affected the cell’s function.
Finally, Proof
Typically, after scientists make changes to DNA, it’s very difficult to prove those changes are actually causing a particular outcome. It’s like seeing lots of switches and lightbulbs turned on, but not being able to know which switch controls a specific light.
In part, that’s because the traditional sequencing approach chops DNA into tiny fragments and reassembles them on the computer, losing key information about any individual molecule.
“Your DNA doesn’t just store information about how to build your body—it also contains a whole control system that decides which parts of those instructions get used, and when,” Ramani says. “Our lab is developing tools to read that control system.”
“This technology is a profound leap that allows us to draw a direct connection that wasn’t possible before.”
—Deepak Srivastava, MD
His team designed a new sequencing method, called SAMOSA, that can read long, intact DNA sequences. In a single readout, his team can see where a stretch of DNA was edited, which genes are turned on, and which parts of the non-coding DNA are active. This means they can directly capture the relationship between the switch and the gene it controls, rather than having to piece it together from separate experiments.
If traditional sequencing was like demolishing every building in a city and trying to figure out which switch was connected to which bulb by looking through the rubble, SAMOSA lets scientists walk into a single building and see a switch connected to a lamp. They can see all the pieces working (or not working) together, in real time.
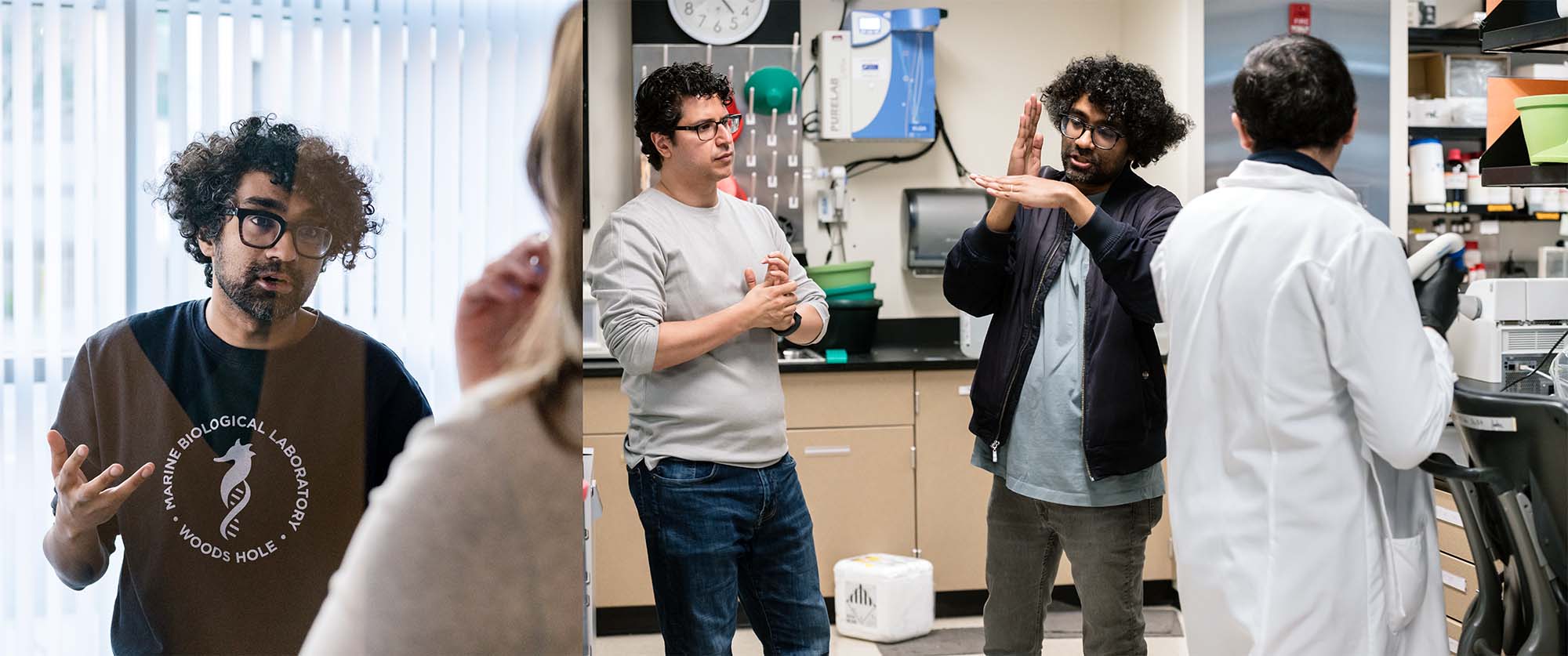

Vijay Ramani (left) and his team developed a DNA sequencing technique that connects non-coding DNA and its influence on a specific gene, allowing scientists to finally understand what changes in the genome are causing disease. (In the photo on the right, Ramani is seen in conversation with his collaborator Hani Goodarzi from Arc Institute (left) and Jose Nunez (right), a research associate in Ramani’s lab.)
“This technology is a profound leap that allows us to draw a direct connection that wasn’t possible before: knowing if a specific stretch of non-coding DNA is active in a particular cell type, and if it’s active, knowing exactly what gene it turns on,” Srivastava says.
With this tool, scientists can now move beyond knowing that a stretch of non-coding DNA is statistically associated with a disease. They can finally see how it functions and influences a specific gene, all in one experiment.
A Growing Circle
Other scientists at Gladstone are also taking part in collaborations that leverage the same iterative process. Christina Theodoris, PhD, brings to the table an AI model called Geneformer, which she developed to predict gene activity within individual cells.
Instead of looking at non-coding DNA, her model can alter a protein-making gene and see how that affects the state of a cell. She and her team then test the predictions—either in her own lab or with Shipman’s group—and then analyze the results, often using Ramani’s measurement tools.
“This allows us to unravel a fundamental puzzle in disease: if a gene is turned on throughout the body, why does a change in this gene only cause disease in, say, the heart, but not everywhere?,” Theodoris says.

Christina Theodoris (center), seen here speaking with Maryam Heidari (right), is developing AI models to predict gene activity within individual cells.
Her work can help target specific genes that, when broken, lead to disease. This will eventually help clinicians know where in the body to screen patients and which organs to target with treatments.
A Loop That Improves the Platform
Once the researchers have gone through this cycle—moving from AI predictions, to genome editing, to a complete readout of the effects—they take everything they learned and go back to the start.
“We’re taking the AI models and supercharging them.”
—Vijay Ramani, PhD
“We’re taking the AI models and supercharging them,” Ramani says. “If a model were trained on all the content from, say, The New York Times, it wouldn’t have much content about heart disease. So, we conduct experiments that model heart disease using genome editing, then we read out the consequences of those edits, and we retrain the model.”
Their goal is to iteratively build the next generation of better AI models by feeding them high-quality data from their experiments, so the models can, in turn, make ever more accurate predictions.
“We’re not just trying to validate an AI model by testing things that are likely to work just to say ‘our model is great,’” Shipman says. “We’re using the model to tell us what experiments are going to be most valuable to make better models. We’re trying to push AI and our molecular technologies as hard as we can so they can eventually give us reliable information where there’s currently uncertainty—as is the case for most diseases.”
Gladstone’s researchers are generating high-quality data from their lab experiments, and using it to iteratively build the next generation of better AI models. (In the photos, graduate student David Wen is working in Theodoris’s lab.)
Solving Together What None Could Solve Alone
One in 31 children in the U.S. has autism—that’s roughly one kid in every other classroom. But of these children, only 20 percent know the specific genetic cause of their condition.
Gladstone’s platform could finally give them a genetic diagnosis. This would guide their medical care, help doctors understand their risk for other health conditions (like seizures), and point to therapies that are most likely to be effective for each person.
I’m confident that we’re going to crack this problem and, for the first time, decode a patient’s DNA so they can alter their genetic destiny.”
—Deepak Srivastava, MD
And the same can be said for many other conditions, from congenital heart defects to Huntington’s disease, where understanding a person’s genome could lead to earlier screening, prevention strategies, and tailored treatments.
“By creating a platform that allows us to decode the language of life as it’s written in DNA, we will ultimately be able to predict disease before it happens and intervene,” Pollard says.
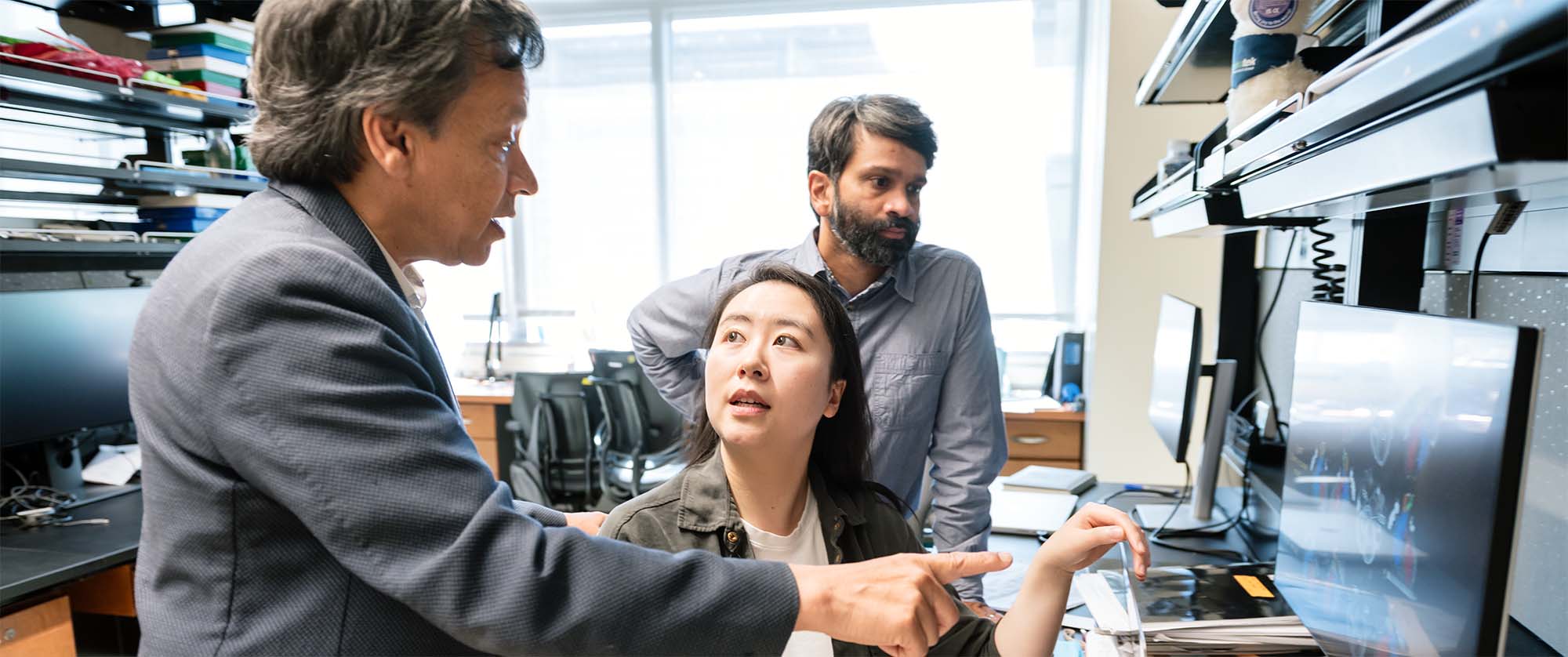

Scientists at Gladstone are bringing together game-changing technologies in a unique way to finally interpret the human genome and find new ways to prevent and treat disease. Seen here is Deepak Srivastava (left) in his lab with Feiya Li (center) and Sanjeev Ranade (right).
Through these efforts, Gladstone scientists are not only finding new ways to prevent and treat disease, but they’re advancing at an unprecedented pace. In the past, from the moment a researcher came up with an initial concept to when that idea could be tested in a human clinical trial would take decades. Now, that timeline can be reduced to fewer than 5 years, Srivastava estimates.
“The technologies that had to come together to understand our genetic code this way don’t exist anywhere else but at Gladstone,” he says. “They were invented here, and we combined them creatively to finally interpret our genomes in ways that have never been possible before. And I’m confident that we’re going to crack this problem and, for the first time, decode a patient’s DNA so they can alter their genetic destiny.”
Gladstone NOW: The Campaign
Join Us On The Journey
Gladstone Launches Center for PhAIge Therapy to Harness AI in the Fight Against Drug-Resistant Infections
Gladstone Launches Center for PhAIge Therapy to Harness AI in the Fight Against Drug-Resistant Infections
The center, funded by an NIH grant, will become one of three national centers dedicated to accelerating the development of phage therapy.
News Release Grants Data Science and Biotechnology Infectious Disease Shipman Lab Pollard Lab Ott Lab Silas Lab AIAI Discovery Reveals DNA Isn’t Locked Away in Cells After All
AI Discovery Reveals DNA Isn’t Locked Away in Cells After All
A new study upends long-held view about how DNA is stored, explaining how subtle shifts in gene activity contribute to cancer, aging, and other complex diseases.
News Release Research (Publication) Cancer Data Science and Biotechnology Ramani Lab Aging AIHow AI Is Pointing Scientists in New Directions and Leading to Novel Drug Targets
How AI Is Pointing Scientists in New Directions and Leading to Novel Drug Targets
Gladstone scientists are developing novel AI models to predict what goes wrong in disease—and how to fix it.
Theodoris Lab AI Cardiovascular Disease



