Gladstone NOW: The Campaign Join Us on the Journey✕

Gladstone Investigator Christina Theodoris is leveraging artificial intelligence to revolutionize how scientists approach crucial questions about the inner workings of the body. She is developing novel models that predict what goes wrong in disease—and how to fix it.
This article is part of a series about the many ways our scientists are using—and developing—AI tools for biomedical research. Sign up for our newsletter to have these stories delivered right to your inbox.
How do brain cells change with age? How are liver cells impacted when someone takes a new medicine? What happens to lung cells after decades of smoking?
The answers to such fundamental questions about cell behavior have direct applications for preventing disease and developing new drugs. And to find the answers, scientists typically design experiments with living cells, tweaking the cells’ DNA or environment to see what happens.
This experimental process has transformed our understanding of human biology and medicine. But it’s also slow, expensive, and difficult to scale up to capture the true complexity of life.
After all, genes that influence cell behavior don’t work in isolation, but through interactions with some of the thousands of other genes in our genomes. Mapping these large genetic networks is an overwhelming or impossible task in the lab, because to fully understand how all genes interact with one another, scientists would need to test an astronomical number of gene combinations.
Now, Gladstone Investigator Christina Theodoris, MD, PhD, is leveraging artificial intelligence (AI) to revolutionize how scientists can approach these crucial questions about the inner workings of the body. She is developing AI models that can predict how cells and their genes will behave under different circumstances—with disease, development, or aging, for instance.
She began by designing Geneformer, the first foundation model in the world to predict how changing gene activity would affect individual cells. This AI model has already revealed new drug targets for heart disease. More recently, she launched another model called MaxToki that can predict what happens to cells throughout the body with aging and how to slow that process.
“When people talk about the power of AI, oftentimes they’re talking about it as a futuristic thing—but it’s already happening today,” says Deepak Srivastava, MD, president of Gladstone. “At Gladstone, we’re developing powerful AI tools to solve challenging diseases that, so far, have been untreatable.”
“When people talk about the power of AI, oftentimes they’re talking about it as a futuristic thing—but it’s already happening today.”
—Deepak Srivastava, MD
In the same way that ChatGPT learned human language by analyzing millions of books and websites, Geneformer and MaxToki learned the language of human genes by analyzing millions of recordings of gene activity across different human cells.
With the new tools, Theodoris and her colleagues can run billions of virtual experiments on the computer, narrowing down the most promising targets to treat a disease before ever stepping into the lab. Rather than the years or decades needed at the lab bench, the new experiments take hours or days.
“By leveraging these computer models to more quickly identify the most promising targets, we can not only accelerate the speed of research, but also move therapies into clinical trials that will have a higher likelihood of success,” Theodoris says.
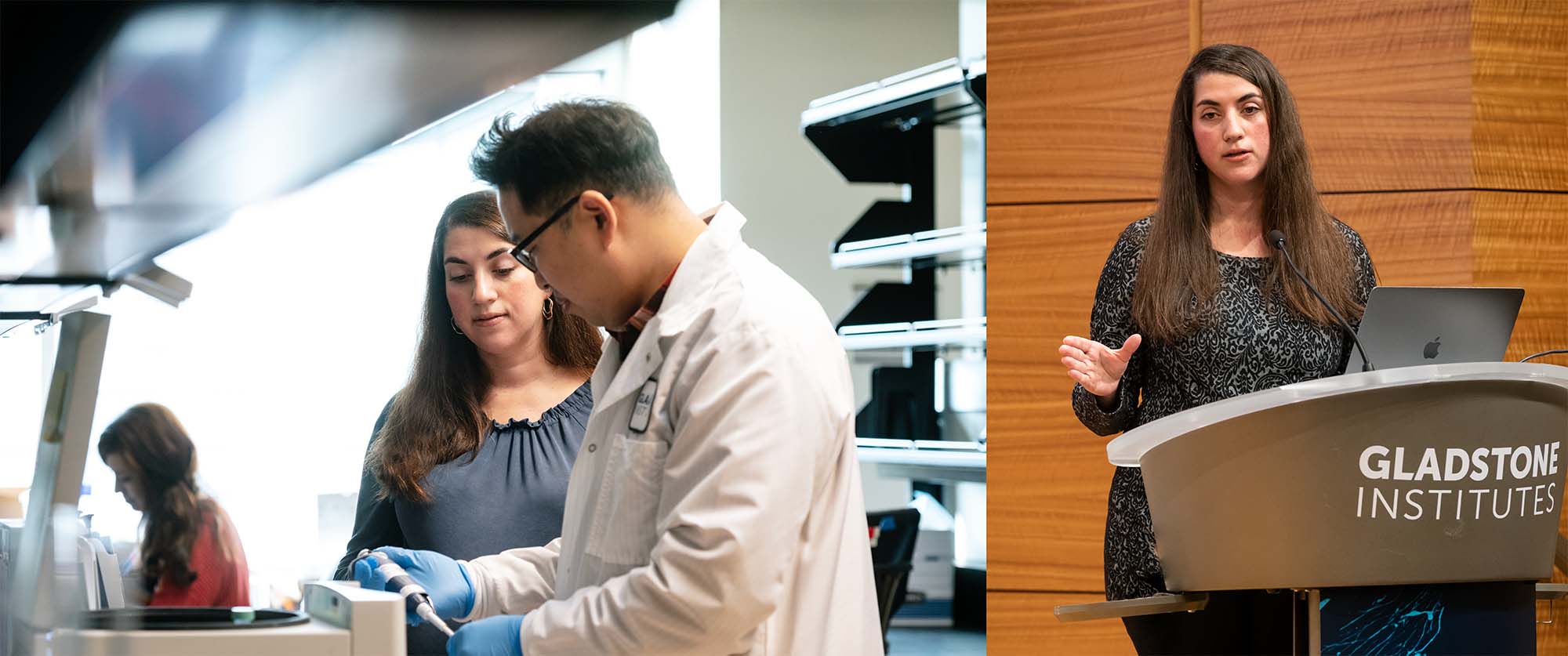

Theodoris (seen here on the left speaking with Bumjoon Kim in the lab) and her team are building large-scale foundation models that are trained on such vast and varied biological data, they develop a fundamental understanding of how cells work. These AI models can then be applied to answer a wide range of questions.
Teaching AI the Language of Cells
Most people are familiar with large language models, like ChatGPT, that have picked up the patterns of human speech and writing. ChatGPT has learned, for instance, that “peanut butter and” is usually followed by “jelly.” So, it can predict that jelly will be the next word when peanut butter is mentioned, and flag “peanut butter and trucks” as an odd combination.
And, just like ChatGPT fundamentally shifted the way internet users search for information, Theodoris’s Geneformer changed how scientists can use AI in their work.
Geneformer is a foundation model, a large-scale AI program trained to recognize patterns across a vast dataset, building general knowledge that can then be applied to many questions. In the same way that ChatGPT learned to associate peanut butter with jelly, Geneformer learned to recognize when certain genes were always turned on at the same time. And, like the mismatch of “peanut butter and trucks,” it can spot patterns of gene activity that are a red flag for disease.
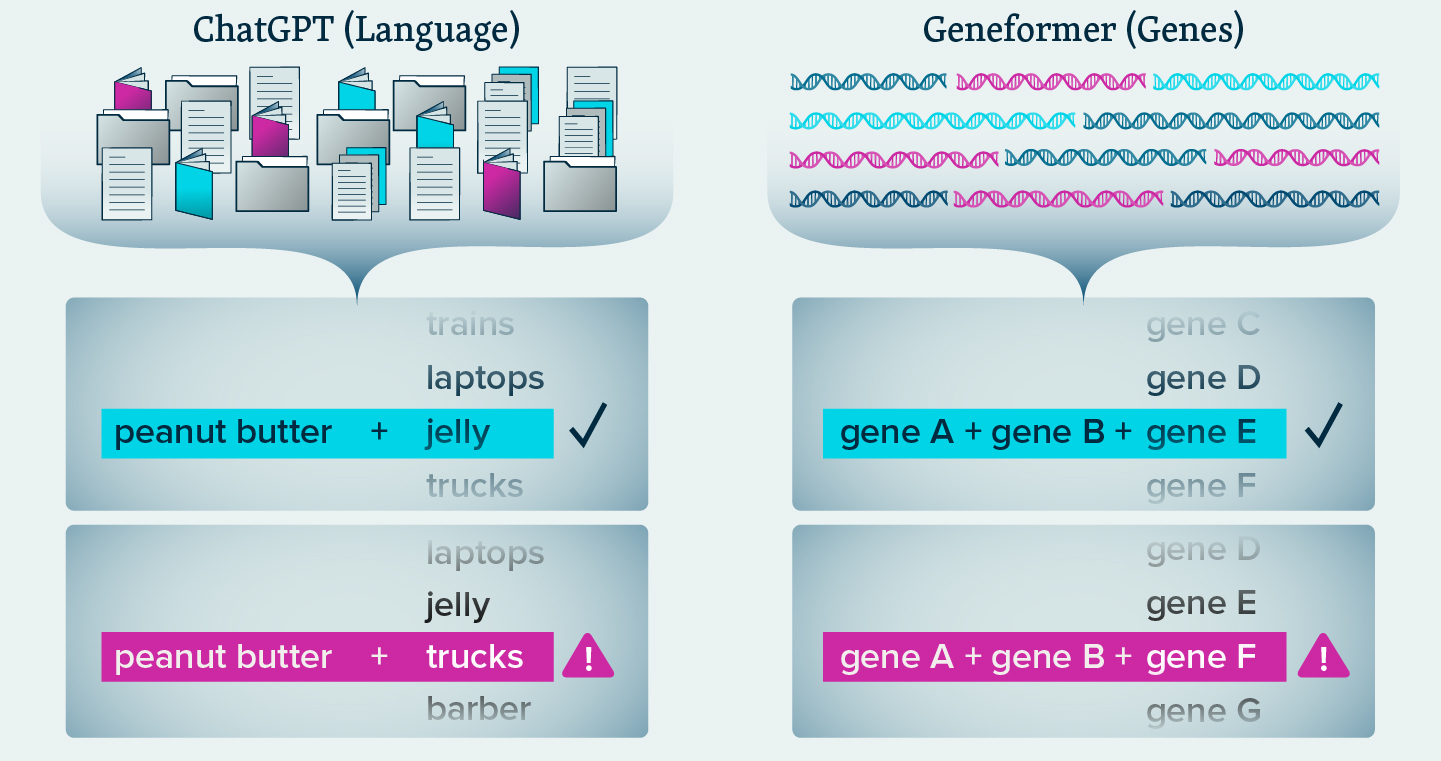
Much like how ChatGPT uses millions of examples of written words to predict and generate new sentences, Geneformer uses information about genes to predict how they interact, and what might go wrong in disease.
“Geneformer learned to recognize which genes to pay attention to in order to predict the levels of other genes, so it can identify the most important genes that control an entire network,” Theodoris explains. “When things go wrong in disease, these are the genes we can target to make cells healthy again.”
Developed in 2021, well before ChatGPT was released, Geneformer was trained initially on 30 million examples of how genes are dialed up and down in individual cells. The data was drawn from public databases and scientific consortia, and spanned a broad range of human tissues, developmental stages, and diseases—essentially everything for which genomic data exists.
It has now been trained on over 100 million examples, and scientists continue to see the model’s predictions improving, which is a promising sign for the future of these approaches as more data becomes available.
“We trained one large-scale model to gain a fundamental understanding of how genes interact in many contexts, and we can now apply it to answer a wide range of questions.” —Christina Theodoris, MD, PhD
The massive scale of the training data, across tissues and contexts, means the model was able to uncover deep relationships in the data and learn general rules about how genes behave across many cell types. It also eliminates the need for researchers to build a new AI tool from scratch for every question about how genes function.
“We trained one large-scale model to gain a fundamental understanding of how genes interact in many contexts, and we can now apply it to answer a wide range of questions,” Theodoris says. “It serves as a general-purpose engine for discovery.”
In addition, Geneformer can make predictions about cells it’s never seen before. This is critical in instances when scientists only have access to limited data, either because the disease they’re studying is rare or it affects tissues that cannot easily be sampled (like the heart or brain).

Theodoris and her team—including Alicja Brozek (left), Abhijay Mahil (center), and Javier Gomez Ortega (right)—are using Geneformer to narrow down the most promising targets to treat a disease on the computer, before ever stepping into the lab.
So, in the same way that you can ask ChatGPT to write a Shakespearean-style sonnet about a food truck—something the model never would have been exposed to—you can ask Geneformer what happens in difficult-to-obtain cells deep in the human body.
“If you want to study these diseases, you wouldn’t have enough data to train a new model, so you need a foundation model with a strong knowledge base to answer your questions,” Theodoris says. “With Geneformer, even for diseases where solutions had been stalled in the past due to limited data, we can finally predict therapeutic targets.”
From Gene Predictions to Potential Treatments
To test the utility of Geneformer, Theodoris and her colleagues used the AI model to study cardiomyocytes, the muscle cells in the heart. The model identified genes that, when disrupted, were most likely to cause problems in the cells.
Many of the genes it listed were already linked to heart disease, indicating it accurately figured out what to look for. But more importantly, the model correctly predicted that these genes matter more in the context of disease—losing them causes more damage than losing most other genes.
Geneformer also predicted genes that had never been studied. And when the researchers removed one of these genes from heart cells in the lab, the cells could no longer beat as robustly.
“It was exciting to us that Geneformer was able to predict a novel key regulator within heart muscle that had never been described before, despite decades of research in these cells,” says Theodoris.
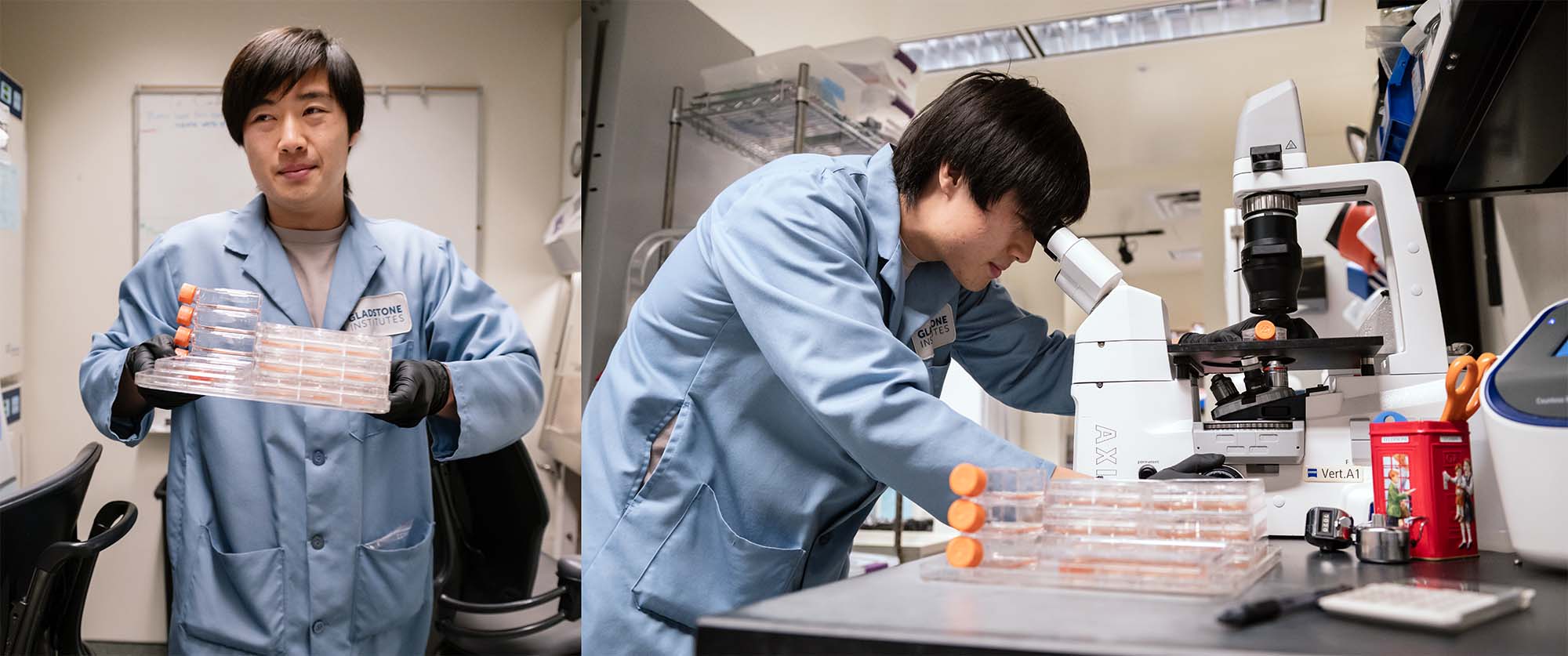

Once the AI model has made predictions, scientists in Theodoris’s group are testing them in the lab. Already, they discovered a potential new therapeutic strategy to treat cardiomyopathy, a disease that affects the heart muscle. (Seen here is David Wen, a graduate student in Theodoris’s lab.)
Next, the team asked Geneformer to predict which genes could be targeted with drugs to restore the function of heart cells in people with cardiomyopathy, a disease of the heart tissue. The AI model homed in on several genes.
In follow-up studies in the lab, Theodoris’s team tested four of those genes in cardiomyocytes. Two of them led to a significant improvement in how strongly the cells contract, and a third showed signs that it was helping the cells beat robustly again—revealing a new therapeutic strategy to treat cardiomyopathy.
“The model was able to point us in new directions to accelerate the discovery of candidate therapeutic targets for this progressive disease,” Theodoris says.
Novel Model to Pinpoint Genes That Drive Aging
Geneformer, for all its power, has a limitation: It sees cells at single points in time. But cells don’t live in static snapshots, they’re dynamic and ever-changing. A neuron in the early stages of Alzheimer’s is different from one late in disease. A heart cell from a 10-year-old is different from one from a 70-year-old.
So, Theodoris built upon the strategy she used for Geneformer to develop a new temporal AI model, MaxToki, which incorporates the dimension of time.
First, her team trained MaxToki—named for a bullet train in Japan whose name is a homonym of the Japanese word for “time”—on data from about 175 million single cells. Then, they assembled 100 million trajectories of cells changing over time, and further trained the model using these cells from thousands of healthy people ranging in age from newborns to more than 90 years old.
The model learned to predict how cells change with aging.

MaxToki, the newest AI model released by Theodoris’s lab, can detect signs of accelerated aging in diseased cells—and predict which malfunctioning genes are driving this aging.
Given an aged cell, MaxToki can reason out which genes changed over time to lead to its final state. And given a diseased cell—which the model had not encountered as part of its training with cell trajectories—it can detect signs of accelerated aging.
The model pinpointed signs of accelerated aging in lung cells from individuals exposed to heavy smoking and patients affected by lung fibrosis. Similarly, in samples from patients with Alzheimer’s disease, the model detected aging acceleration in brain cells. Interestingly, this faster aging was not observed in cells from people whose brains showed signs of Alzheimer’s neuropathology but who had no symptoms of dementia, a phenomenon known as Alzheimer’s resilience.
In addition to recognizing age acceleration in cells affected by diseases of aging, MaxToki could zoom in on exactly how the networks of genes in those cells were going awry, and which malfunctioning genes might be driving the accelerated aging.
In heart muscle cells, the model flagged dozens of genes predicted to either accelerate or slow cardiac aging. The researchers selected five that had never before been linked to aging or disease and tested them in human heart cells grown in the lab.
“To me, the most exciting part of MaxToki is it allowed us to identify novel targets that had a true biological impact on cardiac aging.”
—Christina Theodoris, MD, PhD
When the scientists activated each gene predicted to accelerate aging, the cells showed hallmarks of aging, including irregular beating and dysfunction of genes involved in inflammation and energy usage (by a part of the cell called the mitochondria).
The researchers went on to validate these predictions in a living organism. In fact, when they activated the same genes in young mice, they found a decline in heart function within 6 weeks. The team is now testing whether deactivating the target genes in aged mice can help them be more resilient to the effects of aging.
“To me, the most exciting part of MaxToki is it allowed us to identify novel genes that had a true biological impact on cardiac aging,” Theodoris says. “This could accelerate the discovery of treatments that promote resilience to age-related cardiovascular decline.”
A New Kind of Tool
What makes Geneformer and MaxToki different from previous biology AI tools is their broad applicability. Earlier machine learning models in biology were built for single tasks, such as identifying which cell under a microscope was dividing, or classifying whether a tissue sample contained cancer or not. Every new question meant building a new model, which meant needing enough data to train it.
Foundation models overcome that problem. Because Geneformer and MaxToki were trained on such vast and varied biological data from the start, they developed a broad, generalizable grasp of how cells work. That foundational knowledge can be pointed at new problems without starting over.
“The flexibility of not having to generate a new model for every single question really opens up a lot of possibilities,” says Theodoris. “We can now turn to these models for a wide variety of questions and play out what happens to cells over time when we change gene activity.”

The AI models developed by Theodoris and her team, like Geneformer and MaxToki, are being freely shared with scientists around the world who are asking their own questions about how to control genes to treat disease.
With each disease that Geneformer and MaxToki study, and with each new set of genetic data the models observe, they become better at predicting biology and pointing scientists in new directions.
The AI models developed at Gladstone are now freely shared with research teams around the world who are asking their own questions about how to control genes to treat disease.
“We’re always excited to hear of a new academic lab or pharmaceutical company using our models to predict therapeutic targets for their disease of interest,” Theodoris says. “We want our models to be used widely so they accelerate the discovery of new treatments that benefit patients.”
“Platforms like Geneformer and MaxToki are designed to overcome disease, and to do it faster than we ever could before.”
—Deepak Srivastava, MD
Ultimately, with enough data, Theodoris believes models like Geneformer and MaxToki will uncover the complex rulebook that governs how gene networks are regulated—something that evades biologists today.
“That’s the biggest impact I see for these models in the future,” she says. “It will be tremendous for our ability to understand gene networks more generally, but also for our ability to manipulate these systems to engineer cells or design therapies.”
For patients, that could mean a streamlined path to new drugs. Diseases that have been hard to understand—because the biology was too complex or the cells too hard to access—could be finally solved.
“Platforms like Geneformer and MaxToki are designed to overcome disease, and to do it faster than we ever could before,” says Srivastava. “At Gladstone, we integrate these AI models with our deep disease expertise, and I’m confident our approach will result in cures.”
Gladstone NOW: The Campaign
Join Us On The Journey
This Is AI Predicting New Ways to Treat Disease
This Is AI Predicting New Ways to Treat Disease
How do brain cells change with age? How are liver cells impacted when someone takes a new medicine? What happens to lung cells after decades of smoking?
Cardiovascular Disease Theodoris Lab AINew AI Model Predicts How Cells Age
New AI Model Predicts How Cells Age
Trained on data spanning the human lifespan, a new tool developed at Gladstone can predict genes associated with aging and age-related diseases.
News Release Research (Publication) Cardiovascular Disease Theodoris Lab Aging AIHow AI Is Pinpointing the Genetic Cause of Disease
How AI Is Pinpointing the Genetic Cause of Disease
Scientists at Gladstone are combining AI models with cutting-edge lab tools to finally make sense of the human genome.
AI Srivastava Lab Pollard Lab Engelhardt Lab Theodoris Lab Shipman Lab Ramani Lab Finkbeiner Lab Cardiovascular Disease Neurological Disease



