Gladstone NOW: The Campaign Join Us on the Journey✕
Gladstone scientists have fine-tuned a system for more efficient gene editing, using molecules called retrons.
Over the past decade, the CRISPR genome-editing system has revolutionized molecular biology, giving scientists the ability to alter genes inside living cells for research or medical applications. Now, researchers at Gladstone Institutes have fine-tuned an additional system for more efficient gene editing, using molecules called retrons.
Retrons, the group reported in the journal Nature Chemical Biology, can be optimized for efficiency and used to edit genes in a variety of cell types, from fungi to human cells.
“This work really solidifies retrons as a platform that can be used across organisms,” says Gladstone Assistant Investigator Seth Shipman, PhD, senior author of the new study. “We can make precise modifications to genes more easily, quickly, and efficiently than with current approaches.”
A One-Stop Shop for Gene Editing
Most current gene-editing technologies based on the CRISPR system involve cutting a section of DNA out of a cell’s genome, and then introducing new genetic material called “template DNA” to replace it. As the cell repairs the places where an existing gene was cut, the template DNA is integrated.
That template DNA is normally produced in the lab and then introduced to cells from the outside. The protein that cuts the cell’s genome—called Cas9—is delivered separately. Neither Cas9 nor the template DNA penetrate every cell, limiting the efficiency of CRISPR gene editing.
Retrons, however, act like DNA factories, producing abundant copies of template DNA from inside cells. Moreover, retrons can be delivered along with the rest of the CRISPR components so that cells get all the material needed for gene editing simultaneously—the genetic codes for template DNA, Cas9, and molecules that help researchers track the edits that have been made.
“This means we only have to introduce one element to each cell,” says Santiago Lopez, a graduate student in the Shipman Lab and first author of the new paper. “That significantly simplifies the process and opens the door for new types of experiments.”

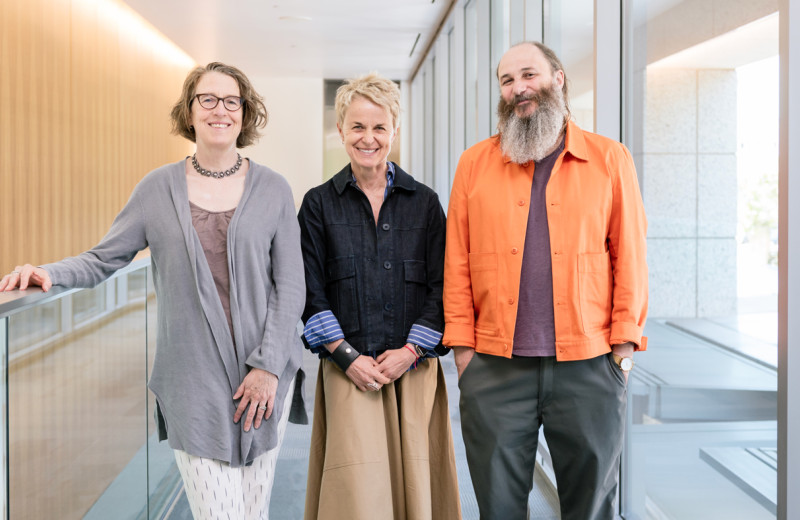
Seth Shipman (left) and Santiago Lopez (right) engineered a new retron system that boosts gene editing efficiency.
Re-Engineering Retrons
Both retrons and CRISPR originate from bacteria; both are defense mechanisms that bacteria use to alter DNA in response to infections. After the advent of CRISPR genome editing, in which the CRISPR system was co-opted to selectively target genes in other cell types, some researchers began probing whether retrons could be used to supply the templates for precise gene editing. However, the roles of different sections of the retron’s structure in its function—and how to tweak those sections to improve retrons—has been unknown.
“The retron system evolved to help defend bacteria,” says Shipman, who is also an assistant professor of bioengineering and therapeutic sciences at UC San Francisco (UCSF). “But we wanted to change it from what it does normally to what we want it to do—produce templates for gene editing.”
In the new study, Shipman’s group engineered E. coli retrons to create hundreds of new variants. They tested each new variant and discovered a series of changes that, together, led to an 8-to-10-fold increase in how much template DNA was eventually produced by the retron in E. coli cells.
Next, the researchers tested the new re-engineered retron system in the fungus Saccharomyces cerevisiae (Baker’s yeast) and in cultured human cells, and they found that this optimized system worked in all cases. This was the first demonstration of retrons’ use in human cells and their portability across cell types.

Seth Shipman (left) and Santiago Lopez (right) developed a useful research tool for editing genes in different cell types in the lab.
Since the team could now fine-tune exactly how much template DNA the retrons produced, they were also able to show that when the retrons produce high levels of template DNA, this boosts gene editing efficiency.
“Our study demonstrates for the first time that the more template DNA we can produce, the better the genome editing,” says Shipman. “Better and more precise editing ultimately means more effective and safer genomic medicines and more advanced fundamental research.”
Taking Tools from Bacteria
Retrons, Shipman says, are immediately useful as a research tool for editing genes in different cell types in the lab. While the platform isn’t yet ready for use in humans, it also holds the potential to help edit genes for therapeutic purposes—for example by repairing gene mutations that cause disease.
Since different bacteria contain different retrons, his group also plans to explore whether other retron variations have benefits over the E. coli retron they optimized in this study.
“We’re taking a general approach in which we’re mining parts that we find in bacteria and domesticating them for our own use,” says Shipman. “This has already been incredibly fruitful for developing new tools, but I think we’re only just beginning to reap the benefits of applying these tools in biotechnology.”
For Media
Julie Langelier
Associate Director, Communications
415.734.5000
Email
About the Study
The paper “Precise genome editing across kingdoms of life using retron-derived DNA” was published in the journal Nature Chemical Biology on December 23, 2021. Other authors are Kate Crawford, Sierra Lear, and Santi Bhattarai-Kline of Gladstone Institutes.
The work was supported by the Simons Foundation Autism Research Initiative, the Pew Scholars Program in the Biomedical Sciences, the Sandler Program for Breakthrough Biomedical Research at UCSF, the National Institutes of Health (1DP2GM140917-01), the L.K. Whittier Foundation, the National Science Foundation (graduate research fellowships 2019247827 and 2034836), a Berkeley Fellowship for Graduate Study, and the Discovery Fellows Program at UCSF.
About Gladstone Institutes
Gladstone Institutes is an independent, nonprofit life science research organization that uses visionary science and technology to overcome disease. Established in 1979, it is located in the epicenter of biomedical and technological innovation, in the Mission Bay neighborhood of San Francisco. Gladstone has created a research model that disrupts how science is done, funds big ideas, and attracts the brightest minds.
Featured Experts
Support Discovery Science
Your gift to Gladstone will allow our researchers to pursue high-quality science, focus on disease, and train the next generation of scientific thought leaders.
Gladstone Launches Center for PhAIge Therapy to Harness AI in the Fight Against Drug-Resistant Infections
Gladstone Launches Center for PhAIge Therapy to Harness AI in the Fight Against Drug-Resistant Infections
The center, funded by an NIH grant, will become one of three national centers dedicated to accelerating the development of phage therapy.
News Release Grants Data Science and Biotechnology Infectious Disease Shipman Lab Pollard Lab Ott Lab Silas LabGladstone Investigator Andrew Yang Receives 2026 Sobrato Prize in Neuroscience
Gladstone Investigator Andrew Yang Receives 2026 Sobrato Prize in Neuroscience
The prize honors scientists whose breakthroughs in brain research have high potential for patient impact.
Philanthropy Awards News Release Alzheimer’s Disease Neurological Disease Yang LabMapping How the Brain Takes Out Its Trash
Mapping How the Brain Takes Out Its Trash
Gladstone scientists developed a new tool to trace how the brain clears “waste,” uncovering surprising new biology about how it keeps itself healthy.
News Release Research (Publication) Alzheimer’s Disease Neurological Disease Yang Lab




