Gladstone NOW: The Campaign Join Us on the Journey✕
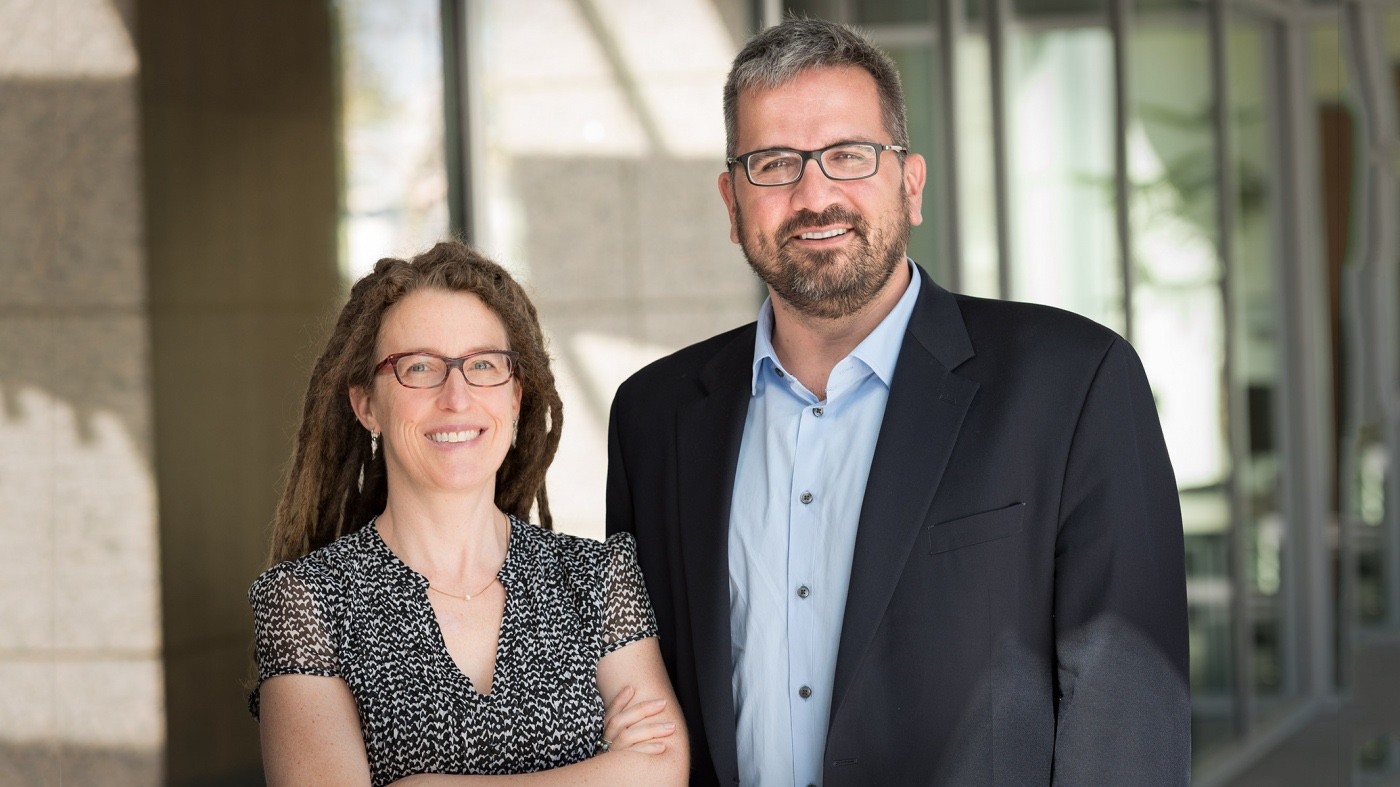

In their pursuit of cures for debilitating diseases, Katherine Pollard, PhD, and Nevan Krogan, PhD, are approaching precision medicine from new angles, researching the microbiome and protein networks that are critical for human biology. [Photo: Chris Goodfellow, Gladstone Institutes]
In 2001, it took 15 months and 300 million dollars to map the 20,000 genes in the human genome. Today, whole genome sequencing technology has become exponentially faster and cheaper. Scientists can now sequence a person’s entire genome in just one day, for only one thousand dollars.
This breakthrough has led to a surge in research on personalized medicine, including cancer treatments that are designed to target specific tumor-causing gene mutations. In fact, scientists have gotten so good at gene sequencing that Nevan Krogan, PhD, a senior investigator at the Gladstone Institutes, is asking his colleagues to shift their focus to the interactions between the genes and the proteins they encode.
“We need to analyze the data we already have more thoroughly in order to understand what these tumor sequences are revealing,” Krogan said.
Beyond DNA
Krogan and his colleagues at Gladstone are researching the genes implicated in various diseases, looking at the proteins they produce and their interactions with other genes in complex networks. Genes rarely act alone. They make proteins that work together with other proteins to perform chemical reactions and to build the infrastructure of cells. Krogan believes that mutations in a few key networks of genes cause diseases like cancer, even though the individual genes that are mutated in tumors vary between different people and different types of cancer. By searching for gene networks with mutations in multiple different genes, his team aims to discover new ways to treat disease.
“If we identify the function of a gene that is mutated in a particular form of cancer, we’re no longer limited to having to develop a targeted therapy for just that one gene. Instead, we can target other genes that impact the same protein or biological process and correct the mutation upstream or downstream in the network,” explained Krogan. “This vastly opens up our options for new druggable targets.”
What’s more, the genes mutated in diseases like cancer or heart disease are often the same genes that are hijacked by invading viruses. For example, Krogan’s lab is working with Jennifer Grandis, MD, at the University of California, San Francisco to investigate the overlap between genes and proteins implicated in head and neck cancer and those attacked by the human papillomavirus (HPV), the number one cause of such cancers. Through this process, the scientists were able to narrow their search and focus on the genes that were both mutated in head and neck cancer and hijacked by HPV.
“We can use viruses like HPV as an entry point to study mutations in head and neck cancer and cervical cancer,” said Krogan. “There may be hundreds of genes mutated in cancer, so identifying the overlapping HPV genes helps us prioritize the really important ones. Studying cancer will help inform research into viral infections and vice versa.”
Krogan is applying this network approach to cardiovascular disease and psychiatric disorders like autism and schizophrenia. He says there are commonalities between the networks that are disrupted in different diseases and also among specific genes. Identifying these cellular Achilles’ heels can lead to similar treatments for seemingly disparate disorders.
Beyond Human Cells
Human biology is not only manipulated by mutations; it is also influenced by the trillions of microbial residents—including viruses—that make up an estimated fifty percent of the cells in the human body.
Gladstone senior investigator Katherine Pollard, PhD, studies the role of this microbial ecosystem on human health and disease. Going beyond the human genome, Pollard advocates for a precision medicine approach at the microbial level, analyzing a person’s microbiome—the genes present within the microbes—to determine the effect it has on conditions varying from autoimmune disorders to body mass to mental health.
“These microbes play an essential role in digestion, metabolism, and the immune system, interacting with our cells in a number of ways,” said Pollard. “If we only study human DNA, we’re going to miss answers to health problems that reside in pathways outside of our genes, which are actually vastly outnumbered by those in the microbiome.”
Human and bacterial cells interact in a number of ways, including physical cell-to-cell contact in which molecules are passed back and forth between the cells. Other bacteria have secretion systems that act like a syringe and directly inject proteins into human cells. These secretions carry important instructions, such as signals that dampen the body’s immune response so that it will not perceive the bacteria as harmful invaders. If these bacteria are eliminated through antibiotics or simple lack of exposure, their immune-dampening response is also lost. This can result in an overactive immune system that attacks the body’s cells, as in the case of autoimmune disorders.
Pollard is investigating this so-called “hygiene hypothesis” to determine which bacterial proteins play a role in immune disorders like inflammatory bowel disease. However, this task is extraordinarily difficult, as the genes within a single species of bacteria can differ by up to fifty percent between strains in different people. And, these genes are not random or trivial; they often have huge implications for disease and are involved in critical processes, like determining which proteins a microbe produces or changing a strain of bacteria from being safe to harmful. Thus, even if two people have the same species of bacteria in their gut, they may be doing completely different things.
So Pollard is trying to pin down not only which bacteria are important for a given disease, but also which proteins—and thus which genes—within that strain of bacteria result in a certain behavior. To date, she has analyzed 120 terabytes of data from more than 2,000 people in 8 countries, with a variety of diseases, in an attempt to identify the bacteria and their genes that affect inflammation and the immune system.
“This type of research wasn’t possible even five years ago,” said Pollard. “Not only do we need the cutting edge technology to identify the bacteria within a given sample, we also need extraordinary computing power to store and analyze all the data. The next step is to gain more information about which genes the microbes are carrying and how they’re interacting with the host genomes.”
Finding Solutions to Disease
Personalized medicine requires more than a single-gene, single-disease approach. Success will come from the integration of the 20,000 human genes, their 100,000 protein products, and the vast microbial ecosystem within the human body. While this is a much more complex and daunting undertaking, by understanding the functions and network interactions of genes and proteins—both human and microbe—we will ultimately gain far greater insight into human health and reveal more solutions to dread diseases.
Support Discovery Science
Your gift to Gladstone will allow our researchers to pursue high-quality science, focus on disease, and train the next generation of scientific thought leaders.
A New Pandemic Threat? Unpacking the Hantavirus Cruise Ship Outbreak
A New Pandemic Threat? Unpacking the Hantavirus Cruise Ship Outbreak
Gladstone virologist Melanie Ott explains why this rare, person-to-person hantavirus strain illustrates the need for diligent pandemic preparedness.
Infectious Disease Krogan Lab Ott LabScientists Map How HIV Hijacks Human Cells—and How Cells Can Fight Back
Scientists Map How HIV Hijacks Human Cells—and How Cells Can Fight Back
A new genetic roadmap reveals hundreds of hidden players in HIV infection, including two proteins that stop HIV in its tracks.
News Release Research (Publication) HIV/AIDS Genomic Immunology Krogan Lab Marson LabCould This Molecule Be ‘Checkmate’ for Coronaviruses?
Could This Molecule Be ‘Checkmate’ for Coronaviruses?
Scientists develop powerful drug candidates that could head off future coronavirus pandemics.
News Release Research (Publication) COVID-19 Infectious Disease Krogan Lab Ott Lab