Gladstone NOW: The Campaign Join Us on the Journey✕
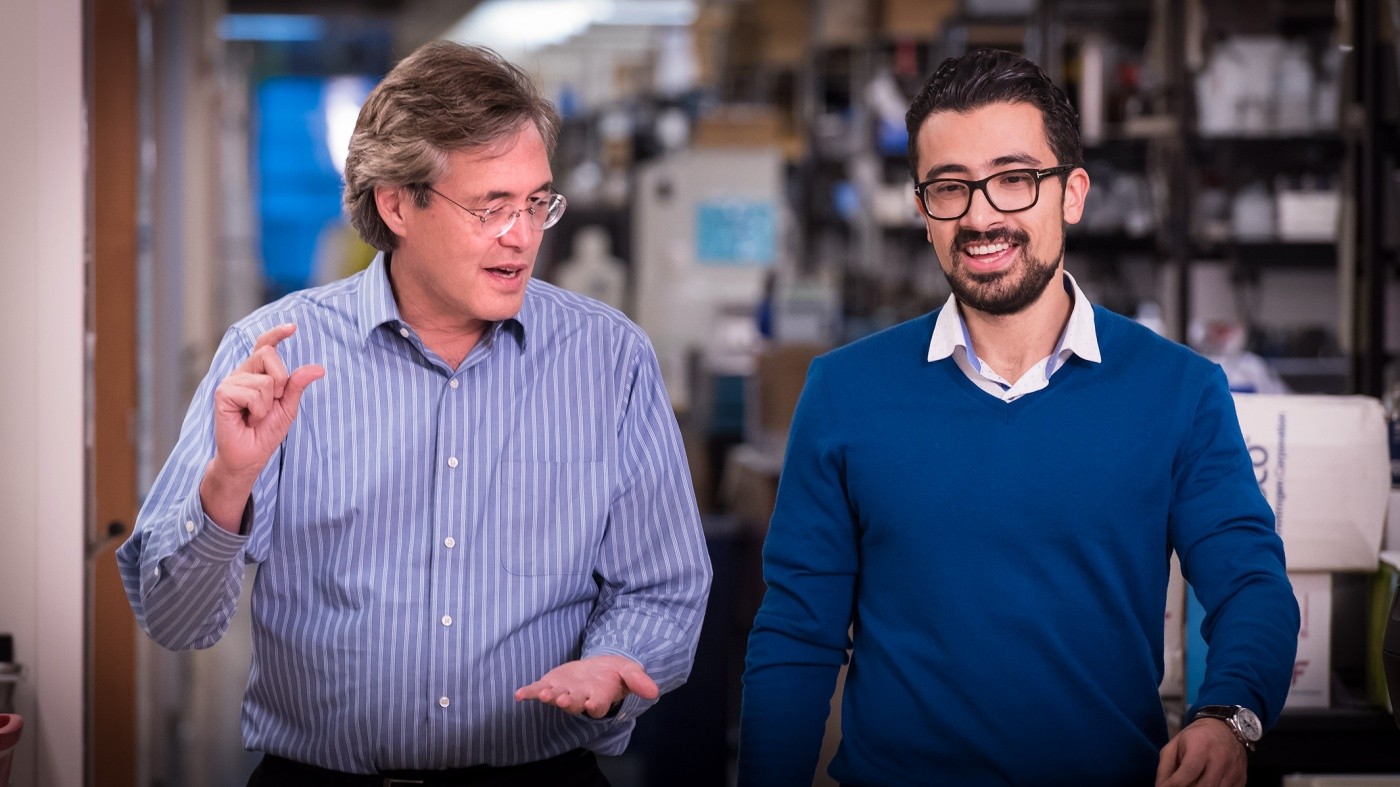

Bruce Conklin, MD, and Mohammad Mandegar, PhD, manipulated the genome of stem cells using a modified form of CRISPR. The new method, called CRISPR interference (CRISPRi) is more precise and efficient than the original CRISPR-Cas9 system, and it offers the flexibility of reversing or controlling the amount of gene suppression. [Photo: Chris Goodfellow, Gladstone Institutes]
Combining the two most powerful biological tools of the 21st century, scientists at the Gladstone Institutes have modified how the genome of induced pluripotent stem cells (iPSCs) is read for the first time using a variation of the CRISPR-Cas9 system. The development offers a major technological advance in creating cell models of genetic diseases.
In a study published in Cell Stem Cell, the researchers used a modified version of CRISPR called CRISPR interference (CRISPRi) to inactivate genes in iPSCs and heart cells created from iPSCs. The method, first reported in 2013 by Stanley Qi, PhD, a co-author on the current paper, significantly improves the original CRISPR-Cas9 system by allowing genes to be silenced—or turned off—more precisely and efficiently. CRISPRi also offers the flexibility of reversing and carefully controlling the amount of gene suppression.
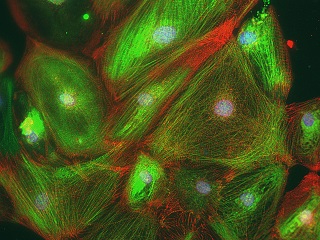 Heart cells created from stem cells, manipulated using CRISPRi. [Image: Mohammad Mandegar, Gladstone Institutes]
Heart cells created from stem cells, manipulated using CRISPRi. [Image: Mohammad Mandegar, Gladstone Institutes]
The standard CRISPR system uses the protein Cas9 to delete a precise part of the genome by making small cuts in a cell’s DNA. CRISPRi builds on this technology by using a special deactivated version of the Cas9 protein and an additional inhibitor protein, KRAB. Together, these proteins sit at the target spot on the genome and suppress gene expression without cutting the DNA. To the scientists’ surprise, temporarily silencing gene expression in this way was much more consistent than permanently cleaving the genome.
“We were amazed by the dramatic difference in performance between the two systems,” said senior author Bruce Conklin, MD, a senior investigator in the Gladstone Institute of Cardiovascular Disease and Roddenberry Stem Cell Center at Gladstone. “We thought that permanently cutting the genome would be the more effective way to silence a gene, but in fact, CRISPRi is so precise and binds so tightly to the genome that it is actually a better way to silence a gene.”
In the study, the researchers compared how well CRISPRi and CRISPR-Cas9 silenced a particular gene that controls iPSC pluripotency—the capability of iPSCs to turn into multiple cell types. They discovered that CRISPRi was much more efficient than CRISPR-Cas9: in more than 95% of the cells created using CRISPRi, the target gene was silenced, compared with only 60-70% of cells grown from CRISPR-Cas9. CRISPRi also did not cause any off-target changes in gene expression, like undesired insertions or deletions to the cell’s genome, which is a concern with CRISPR-Cas9.
CRISPRi also acted like a toggle switch, enabling the scientists to reverse gene suppression by simply removing the chemical that turns on the gene inhibitor. In addition, the researchers were able to tune how much they silenced a gene by changing the amount of the chemicals they added. Both of these results support much more versatile investigations into the role of certain genes that affect development and disease.
The researchers demonstrated the enhanced capabilities of CRISPRi in iPSCs, T cells, and heart cells made from stem cells. For example, they created a model of heart disease using genome editing to suppress a gene essential for heart function.
“CRISPRi holds a major advantage in making disease-relevant cell types,” said first author Mohammad Mandegar, DPhil, a research scientist at Gladstone. “Using this technology, we can mimic disease in a homogenous population of heart cells created from iPSCs. This development allows us to study genetic diseases more easily and potentially identify new therapeutic targets.”
Gene Editing Strategy Could Treat Hundreds of Inherited Diseases More Effectively
Gene Editing Strategy Could Treat Hundreds of Inherited Diseases More Effectively
Scientists at Gladstone show the new method could treat the majority of patients with Charcot-Marie-Tooth disease.
News Release Research (Publication) Neurological Disease Conklin Lab CRISPR/Gene EditingGladstone’s Scientific Highlights of 2025
Gladstone’s Scientific Highlights of 2025
From fundamental insights to translational advances, here’s how Gladstone researchers moved science forward in 2025.
Gladstone Experts Alzheimer’s Disease Autoimmune Diseases COVID-19 Neurological Disease Genomic Immunology Cardiovascular Disease Data Science and Biotechnology Infectious Disease Conklin LabThe Genome Editing Playbook Is Different in Neurons
The Genome Editing Playbook Is Different in Neurons
The striking findings of a new study could influence how gene therapies are designed for many genetic diseases.
News Release Research (Publication) Neurological Disease Conklin Lab Doudna Lab CRISPR/Gene Editing