Gladstone NOW: The Campaign Join Us on the Journey✕
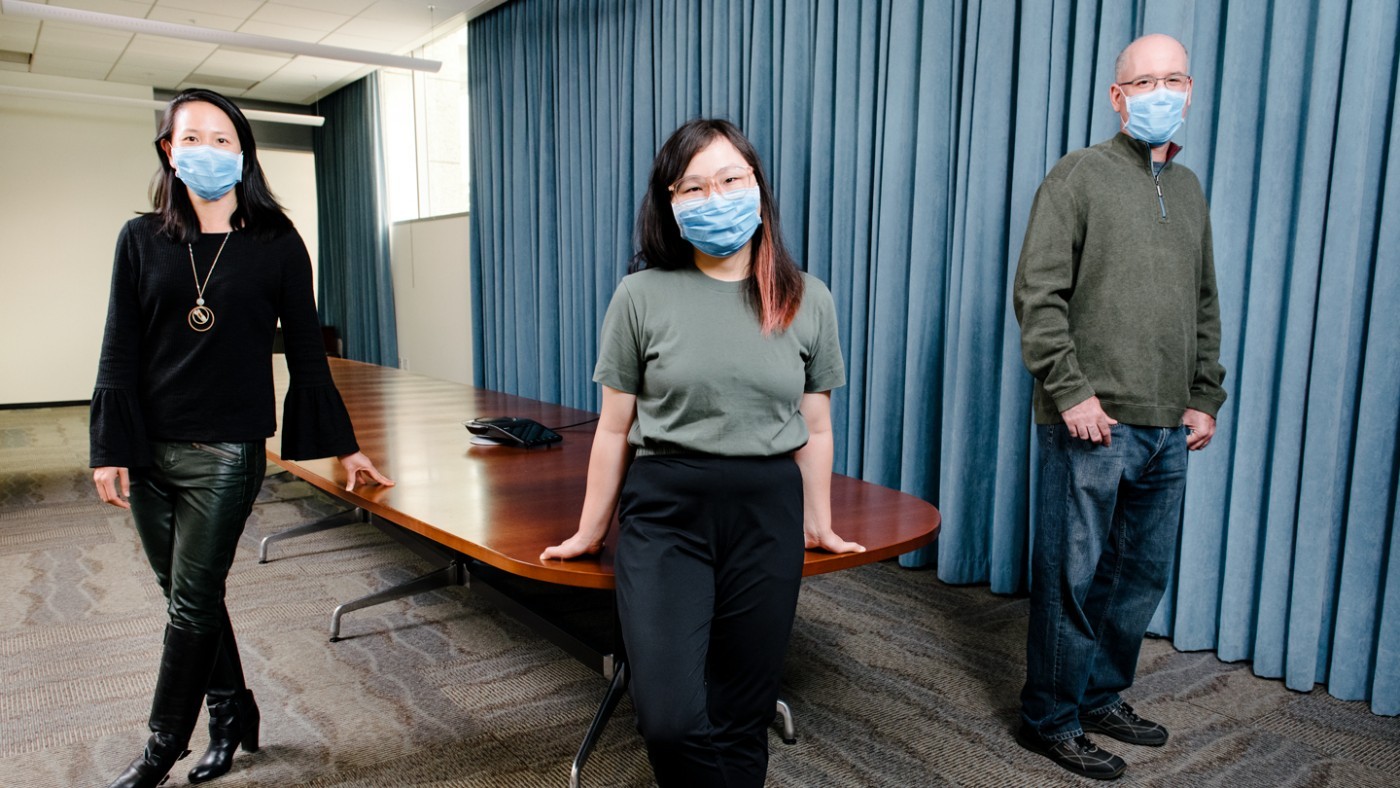

Nadia Roan (left), Xiaoyu Luo (center), and Jason Neidleman (right) mapped out an atlas of latent reservoir cells that could open new avenues for studying or targeting the main barrier to an HIV cure.
The latent reservoir is the last bastion of HIV’s resistance to a cure. But it is difficult to destroy because it is invisible: the cells in the reservoir harbor virus that is dormant, so they don’t have any viral proteins on their surface that would give them away.
As a result, scientists have struggled to learn what the reservoir looks like in individuals with HIV. And without this knowledge, they harbor little hope of being able to target the reservoir with therapies that could eliminate or reduce it, thus ridding people of HIV infection for good.
To fish out reservoir cells, scientists have to reawaken the virus by activating cells they collect from infected individuals. Once awake, the virus produces proteins that mark the surface of its host cells, which gives researchers a handle to find and study these cells. However, the very process of reactivating the virus leads to changes to the cells’ biology that obscure their original identity. And so, the true identity of the cells making up the latent reservoir—also called latent cells—has remained elusive.
To overcome this problem, Gladstone Scientist Nadia Roan, PhD, has taken advantage of an approach she developed previously to backtrack reactivated reservoir cells to their original latent state. With this approach, Roan and her team have mapped out an atlas of the reservoir cells of eight individuals living with HIV, which they recently reported in the journal eLIFE.
“Our findings challenge some previously held assumptions about the makeup of the reservoir,” says Roan, who is also an associate professor of urology at UC San Francisco. “In addition, our detailed map of reservoir cells will make it easier to find these cells in infected individuals, which will fundamentally change how the latent reservoir can be studied.”
Method to the Madness
Previous investigations suggest that the reservoir consists in large part of memory T cells, a subset of cells in the immune system that retain the memory of past infections. These cells can remain for a long time in the body in a quiescent state, waiting for a new infection by a previously encountered virus to wake them up. That makes them the perfect hiding place for HIV.
But memory T cells come in many types, and a commonly held view among scientists is that the latent reservoir consists of a random assortment of memory T cells rather than a specific subset.
“If so, targeting latently infected cells as a curative strategy would be all the more difficult,” says Roan.
Most previous studies of the reservoir have relied on the examination of only a few proteins on the surface of cells. With Roan’s approach, however, her team can follow nearly 40 proteins at once, which greatly increases their ability to distinguish even closely related cells. The researchers can also compare populations of cells before and after reactivation, thus matching each reactivated cell to the pre-activation cell that resembles it most, as if going back in time.
“It’s somewhat like implementing facial recognition technology on cells,” says Roan. “You can think of it as having a photo of someone in their fifties, and trying to identify them in their high school yearbook. Although individuals’ looks change as they age, you can typically still recognize them by looking for a combination of their traits. Similarly, latent cells change as they are reactivated, but they still retain some of their original identity in a way that we can capture by tracking 40 proteins at once.”
Roan’s team carried out this analysis on millions of cells collected from eight individuals under antiretroviral therapy. The cells came from the donors’ blood and gut, the latter which is thought to be a primary site of viral persistence in individuals on this treatment. The scientists also obtained cells from the lymph nodes of one donor.
“Reservoir cells reside in the blood but also in various tissues in the body,” says Jason Neidleman, a senior research associate in Roan’s lab and co-first author of the study. “We wanted to know how reservoir cells in the blood compare to those from other sites.”
By analyzing millions of cells patient samples, Jason Neidleman, the study's co-first author, helped build an atlas of cells that HIV can infect.
The team first built an atlas of the CD4+ T cells (the type of T cells HIV can infect) in each infected donor, based on the assortment of the 40 proteins these cells contained. Then, they mapped each reactivated cell from each individual against the corresponding atlas to find the most similar atlas cell. This most similar cell is thought to represent the original state of the latently infected cell, before it reawakened from latency.
“Somewhat to our surprise, we found that the blood reservoir is not randomly distributed among memory T cells,” says Xiaoyu Luo, PhD, a scientist at Gladstone and co-first author of the study. “Instead, reservoir cells in the blood samples map to a few distinct areas on the atlas. What’s more, reservoir cells from different donors mapped near one another, indicating that they share common features.”
When comparing the blood, lymph nodes, and gut samples, the team also revealed important differences between blood and tissue cells, but also some shared markers, in particular between gut and lymph node cells.
“The existence of shared features across people and tissue types give us hope that we can one day design therapies that target large fractions of the reservoir at once and will work for many infected individuals,” says Roan.
Toward a Deeper Understanding of the Reservoir
For now, the team is keen to use their findings to learn more about the reservoir.
“One of the problems that has galled the field is that most reservoir cells harbor defective versions of the HIV genome,” says Roan. “These cells do not constitute the most clinically relevant reservoir of HIV, because even after they are reactivated, they do not produce infectious virus.”
This situation makes it difficult to home in on the reservoir cells that truly matter—those with a viral genome competent for replication and infection—which can represent as little as 1 percent of the reservoir population.
However, Roan and her team found that when they used the shared markers they identified to extract cells from donor samples, they could obtain populations of cells where more than 50 percent of the reservoir cells contained intact viral genomes.

Along with other members of Roan's team, co-first author of the study Xiaoyu Luo found shared features across cells from different individuals with HIV, which could help develop novel therapies.
“These results suggest that intact and defective virus are kept in different subsets of cells,” says Roan. “And now that we can more readily identify the reservoir cells capable of producing infectious virus, we can begin to elucidate how these cells persist in an infected individual over time.”
Another problem of infectious reservoir cells is that they are very rare to begin with—perhaps as few as one in a million CD4 T cells—which makes it all the more difficult to obtain enough replication-competent reservoir cells to carry out experiments. The markers Roan’s team identified alleviate this problem by allowing researchers to increase the proportion of infectious reservoir cells in a donor’s sample by 100-fold or more.
“By increasing our access to infectious reservoir cells this much, we open up the possibility of conducting a variety of previously impossible experiments that could greatly refine our understanding of reservoir cells,” says Roan. “In particular, it might allow for the discovery of unanticipated—and perhaps even unique—markers of latent cells, which could speed up the design of new therapies for HIV eradication.”
For Media
Julie Langelier
Associate Director, Communications
415.734.5000
Email
About the Study
The paper "Phenotypic analysis of the unstimulated in vivo HIV CD4 T cell reservoir" was published in eLife on September 29, 2020.
Other researchers on this study include Julie Frouard, Grace Xie, Feng Hsiao, Tongcui Ma, Reuben Thomas, and Warner Greene from Gladstone Institutes; Vincent Morcilla, Ashley Lee, and Sarah Palmer from the University of Sydney, Australia; Sushama Telwatte and Steven A. Yukl from San Francisco Veterans Affairs Medical Center; Whitney Tamaki, Benjamin Wheeler, Rebecca Hoh, Peter W. Hunt, Steven G. Deeks, Ma Somsouk, Poonam Vohra, and Jeffrey Milush from UC San Francisco; and Katherine Sholtis James and Nancie M. Archin from the University of North Carolina at Chapel Hill, North Carolina.
This study was supported by the National Institutes of Health (R01AI127219 and R01AI147777, R01AI134363, UM1AI126611, P01AI131374), the amfAR Institute for HIV Cure Research, and the Pendleton Foundation.
About Gladstone Institutes
Gladstone Institutes is an independent, nonprofit life science research organization that uses visionary science and technology to overcome disease. Established in 1979, it is located in the epicenter of biomedical and technological innovation, in the Mission Bay neighborhood of San Francisco. Gladstone has created a research model that disrupts how science is done, funds big ideas, and attracts the brightest minds.
Featured Experts
Support Discovery Science
Your gift to Gladstone will allow our researchers to pursue high-quality science, focus on disease, and train the next generation of scientific thought leaders.
A Gene That Keeps Stem Cells From Losing Their Way
A Gene That Keeps Stem Cells From Losing Their Way
Scientists find a gene essential for keeping intestinal stem cells stable, offering insight into how tissues repair themselves.
News Release Research (Publication) Yamanaka Lab Stem Cells/iPSCsGladstone Recognized in Inaugural Cure Innovation Index for Translating Science Into Real-World Health Impact
Gladstone Recognized in Inaugural Cure Innovation Index for Translating Science Into Real-World Health Impact
Gladstone ranks 15th among 60 institutes and centers nationally for its ability to turn discoveries into measurable health outcomes.
Institutional News News ReleaseAI Discovery Reveals DNA Isn’t Locked Away in Cells After All
AI Discovery Reveals DNA Isn’t Locked Away in Cells After All
A new study upends long-held view about how DNA is stored, explaining how subtle shifts in gene activity contribute to cancer, aging, and other complex diseases.
News Release Research (Publication) Cancer Data Science and Biotechnology Ramani Lab Aging AI




