Gladstone NOW: The Campaign Join Us on the Journey✕
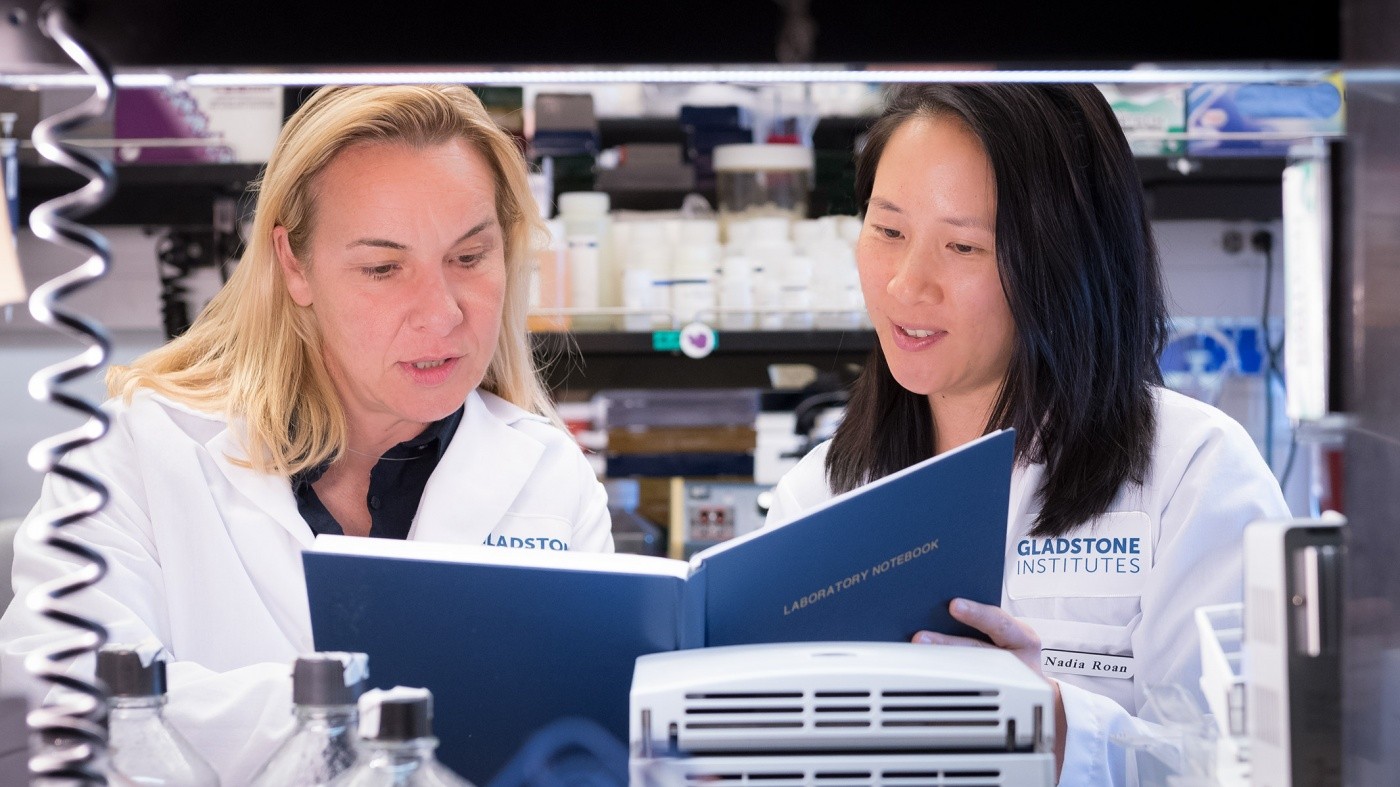

Nadia Roan (right) and Marielle Cavrois (left) identified the cells most likely to be infected by HIV. [Photo: Chris Goodfellow, Gladstone Institutes]
HIV primarily targets a type of cell in the immune system called the CD4+ T cell. However, many types of these cells exist, and they are not all equally susceptible to infection by HIV. But exactly which types of cells are most likely to be infected?
A team of scientists at the Gladstone Institutes and the University of California, San Francisco (UCSF) created a “cellular atlas” to answer this question. They are the first group to globally analyze the different types of CD4+ T cells that HIV enters and manipulates.
“Using powerful new technologies, we identified which cells HIV likes to enter and infect, and how the virus is able to alter the cells once inside them,” explained Nadia R. Roan, PhD, lead author of the study published in Cell Reports and a visiting investigator at Gladstone. “The changes made to cells by the virus can make it easier for HIV to persist in the body.”
In collaboration with Stanford, the University of Southern California (USC), and the University of Alabama (UAB), the team used experimental, statistical, and visualization approaches to create the atlas.
The main tool they used is called CyTOF, which relies on mass spectrometry to measure many parameters on individual cells. This high-resolution technology surpasses other commonly used techniques that limit the number of parameters that can be detected at one time.
“With CyTOF technology, we are entering an era of ‘facial recognition’ for individual cells,” said Marielle Cavrois, first author of the study, a senior staff research scientist in Gladstone’s Flow Cytometry core, and an assistant professor in the Department of Medicine at UCSF. “With our computer tools, we can now use this facial recognition to determine what the cell looked like before HIV manipulated it.”
Given that HIV dramatically changes the cell it infects, it is difficult to trace an infected cell back to its initial form. With CyTOF, the researchers were able to study infected cells in such great detail that they could monitor how HIV remodeled them.
“By comparing the cells that are susceptible to HIV entry to those susceptible to HIV infection, we made a surprising discovery,” added Roan, who is also an assistant professor in the Department of Urology at UCSF. “We found that HIV enters a specific type of CD4+ T cell very efficiently, but is not able to multiply itself in these cells.”
This effect could be because HIV can’t progress any further in those cells, or because the virus remains silent (or latent) inside them. Either way, this subset of CD4+ T cells could become a target for eradicating the virus.
“It will be interesting to discern what is happening in these cells where HIV enters efficiently but does not go on to produce virus,” said Warner C. Greene, senior investigator and director of the Gladstone Institute of Virology and Immunology, who is also one of the senior authors of the study. “If indeed the virus is establishing a latent infection in these cells, we will have identified a valuable biomarker that could lead to new therapies.”
Other authors of this resource article include Nandhini Raman at Gladstone; Rajaa Hussien, Brandon Aguilar Rodriguez, Joshua Vasquez, Matthew H. Spitzer, and Joseph M. McCune at UCSF; Trambak Banerjee and Gourab Mukherjee at USC; Nicole H. Lazarus, Eugene C. Butcher, Ann M. Arvin, and Nandini Sen at Stanford; and Jennifer J. Jones and Christina Ochsenbauer at the UAB at Birmingham. This work was supported by the National Institutes of Health (R01AI127219, DP1DA036502) and the amfAR Institute for HIV Cure Research.
Want to Join the Team?
Our people are our most important asset. We offer a wide array of career opportunities both in our administrative offices and in our labs.
Explore CareersScientists Map How HIV Hijacks Human Cells—and How Cells Can Fight Back
Scientists Map How HIV Hijacks Human Cells—and How Cells Can Fight Back
A new genetic roadmap reveals hundreds of hidden players in HIV infection, including two proteins that stop HIV in its tracks.
News Release Research (Publication) HIV/AIDS Genomic Immunology Krogan Lab Marson LabScience in Seconds | A New Path Toward Life Without Daily HIV Pills
Science in Seconds | A New Path Toward Life Without Daily HIV Pills
In this video, Nadia Roan and Ashley George explain how they uncovered a new path toward long-term health without the need for daily HIV pills.
Gladstone Experts Research (Publication) HIV/AIDS Infectious Disease Roan LabWhy Some People Naturally Control HIV Even After Stopping Therapy—and How We Can Leverage That to Treat Others
Why Some People Naturally Control HIV Even After Stopping Therapy—and How We Can Leverage That to Treat Others
New research offers a path toward life without daily HIV pills, suggesting common diabetes pill could help achieve long-term remission.
News Release Research (Publication) HIV/AIDS Infectious Disease Roan Lab