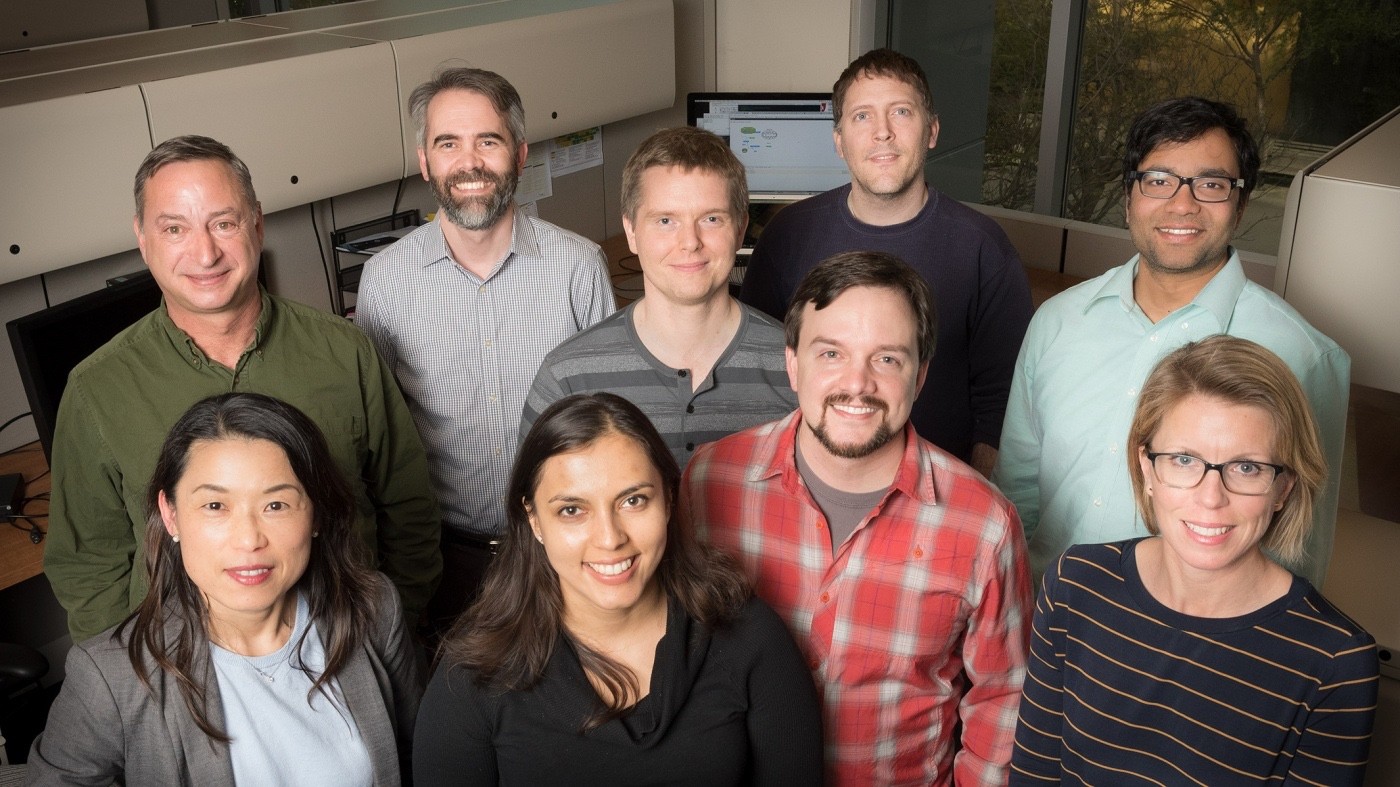
The Gladstone Bioinformatics Core provides expertise in data analysis and visualization to answer complex biological questions and push discoveries closer to cures. [Photo: Chris Goodfellow, Gladstone Institutes]
Technology has rapidly accelerated in the sciences, creating a new era of discovery. Scientists are now awash in data, and the challenge is often how to make sense of all the information. To keep up with the influx of Big Data, the Gladstone Institutes has a dedicated Bioinformatics Core. These scientists provide expert data analysis and visualization, pushing discoveries closer to cures.
The Core works with laboratories across Gladstone to answer complex biological questions and provide insights into a wide range of diseases. The nine members of the bioinformatics team, led by Core Director Eva Wang, PhD, have expertise in statistics, computer science, biology, and engineering. Using complex analyses and major computing power, they collaborate with their peers to dive deep into a single cell to uncover the integrated networks of proteins and genes that lie within, or they scan through millions of variations in human DNA to identify the ones that cause disease.
“Quantitative biology permeates all aspects of research at Gladstone,” said Katherine Pollard, PhD, director of Gladstone’s convergence science initiative who founded the Bioinformatics Core when she joined Gladstone in 2008. “Regardless of the topic they’re working on, everyone has big, complex data, and they need experts to get the most out of that data.”
Family Genome Sequencing Provides New Insights into Neurodegenerative Disease
One of the Core’s long-standing projects is a collaboration with the laboratory of Steven Finkbeiner, MD, PhD, a director at Gladstone. Together, they are probing the genomes of families with Huntington’s disease.
Huntington’s disease stems from a gene mutation that causes the huntingtin protein to misfold and accumulate in neurons, severely damaging the brain. However, family members who carry the same disease-causing gene mutation can develop Huntington’s at vastly different ages. For example, one sibling may show signs of the disease in their 30s, while another may not develop symptoms until well into their 70s.
“We think the differences in age of onset between family members are due to differences in other genes. Our question is, what are those other genes, what functions do they have, and how do they modulate disease onset?” explained Julia Kaye, PhD, a program manager and staff research scientist in the Finkbeiner laboratory. “We hope to find these genetic modifiers of Huntington’s disease by looking at the inheritance patterns of genetic variants from whole genome sequence data in families.”
Kaye is working with members of the Bioinformatics Core to analyze the genome sequence data of nearly 150 people from 23 families with Huntington’s disease. Using complicated computational and statistical pipelines, the researchers narrow down the millions of genomic variants that exist between the patients to a few thousand relevant contenders that may influence the age of disease onset.
So far, Kaye is excited about one particular variation that affects the expression of a gene implicated in the breakdown of proteins in a cell. She thinks this gene may have enhanced expression in people with late-onset Huntington’s disease, which could help prevent the huntingtin protein from accumulating in the brain. If she is right, the gene could be a viable therapeutic target to treat or delay the onset of Huntington’s disease.
Single-Cell Analysis Reveals Secrets of the Heart
The Core is also working with Senior Investigator Benoit Bruneau, PhD, to characterize cells important for heart development. Using an innovative new technique called single-cell RNA sequencing, they aim to learn exactly which proteins a cell produces at a given time and thus which genes are active.
Earlier methods for sequencing RNA were applied to chunks of tissue, which contain dozens of different cell types that had to be sampled together. The analyses measured the average RNA expression of all the cells, which told the scientists the most prevalent genes that were active in the tissue, but not those that were active in only a few cells. Scientists have been longing to capture these minority cells, because they are often the drivers of tissue development and disease.
In the new approach, the scientists can sequence RNA in individual cells, allowing them to characterize all the different types of cells in a given sample. However, because this method sequences thousands of cells individually instead of en masse, it produces exponentially more data.
Thanks to analytical help from members of the Core, Bruneau’s team is now using single-cell RNA sequencing to gain deeper insights into how many of each type of cell are present during the different stages of heart development and what they are doing. The researchers are also applying the method to learn more about the differences between healthy and diseased hearts, identifying all the changes in gene expression that result from a single mutation.
“As biologists, we know the questions that we want to ask of the data, but we don’t have the practical or theoretical knowledge of how to ask those questions,” said Bruneau. “We need statisticians, computer scientists, and bioinformaticians to formulate the analytics and then carry them out.”
Full-Service Science
One of the Core’s strengths is that they are fully integrated in the research process from start to finish. In addition to analyzing data, the bioinformatics team consults on study inception and design, and they help generate figures for grants and scientific manuscripts.
Wang commented, “A longer-term collaborative process is optimal for both sides, because it helps us understand the data better, and it helps the labs design the studies more effectively to get more information out of their data.”
Wang is also expanding the scope of the Bioinformatics Core to support scientific research beyond Gladstone. One example is the work of Core Associate Director Alex Pico, PhD, on open source software that helps scientists to model and visualize their data more easily. Two programs his group develops, WikiPathways and Cytoscape, allow researchers to place genes and proteins into interactive pathways and networks so that they can better understand the molecules’ functions and interactions. These programs have been used in tens of thousands of scientific publications, contributing to science around the world.
Whether through collaborations at Gladstone or through the development of advanced resources available to all, the Bioinformatics Core helps researchers dig deeper into their data to make intricate and multifaceted connections between genes, proteins, and cells. By doing so, they facilitate scientists in their pursuit of knowledge and, ultimately, their quest for cures.
Want to Join the Team?
Our people are our most important asset. We offer a wide array of career opportunities both in our administrative offices and in our labs.
Explore CareersConnecting the Dots: From Stem Cells to Human Health
Connecting the Dots: From Stem Cells to Human Health
Since Shinya Yamanaka discovered a method to create human induced pluripotent stem (iPS) cells, Gladstone researchers have built on this technology to create numerous iPS cell lines, coax iPS cells into a myriad of tissue types, use them as a tool to study disease, and even begin creating therapies using these cells.
Research (Publication) Stem Cell Core Huntington’s Disease Alzheimer’s Disease Finkbeiner Lab Huang Lab McDevitt Lab Srivastava Lab Yamanaka Lab CRISPR/Gene EditingTaube Philanthropies Announces $3 Million Gift to Stanford Medicine and Gladstone Institutes for Huntington’s Disease Research
Taube Philanthropies Announces $3 Million Gift to Stanford Medicine and Gladstone Institutes for Huntington’s Disease Research
The funded program will, for the first time, introduce gene editing and stem cell therapies in the search for a potential cure
Huntington’s Disease Finkbeiner LabDisease Models Facilitate Breakthrough Science
Disease Models Facilitate Breakthrough Science
Scientists have developed sophisticated models to unravel complex human biology in their pursuit of cures.
Finkbeiner Lab McDevitt Lab Ott Lab Sanjabi Lab Zika Virus Hepatitis C Huntington’s Disease